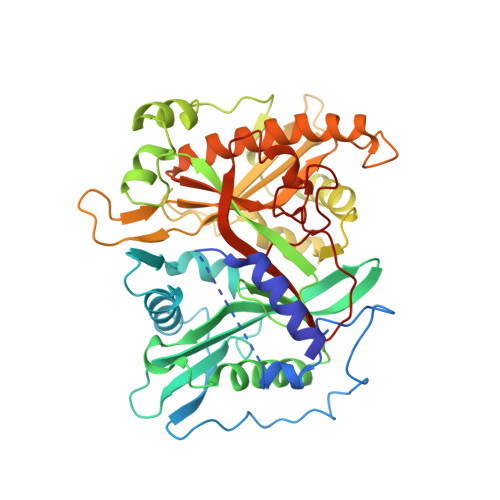
Crystal structures of Saccharomyces cerevisiae N-myristoyltransferase with bound myristoyl-CoA and inhibitors reveal the functional roles of the N-terminal region.
Wu, J., Tao, Y., Zhang, M., Howard, M.H., Gutteridge, S., Ding, J.(2007) J Biol Chem 282: 22185-22194
- PubMed: 17513302
- DOI: https://doi.org/10.1074/jbc.M702696200
- Primary Citation of Related Structures:
2P6E, 2P6F, 2P6G - PubMed Abstract:
Protein N-myristoylation catalyzed by myristoyl-CoA:protein N-myristoyltransferase (NMT) plays an important role in a variety of critical cellular processes and thus is an attractive target for development of antifungal drugs. We report here three crystal structures of Saccharomyces cerevisiae NMT: in binary complex with myristoyl-CoA (MYA) alone and in two ternary complexes involving MYA and two different non-peptidic inhibitors. In all three structures, the majority of the N-terminal region, absent in all previously reported structures, forms a well defined motif that is located in the vicinity of the peptide substrate-binding site and is involved in the binding of MYA. The Ab loop, which might be involved in substrate recognition, adopts an open conformation, whereas a loop of the N-terminal region (residues 22-24) that covers the top of the substrate-binding site is in the position occupied by the Ab loop when in the closed conformation. Structural comparisons with other NMTs, together with mutagenesis data, suggest that the N-terminal region of NMT plays an important role in the binding of both MYA and peptide substrate, but not in subsequent steps of the catalytic mechanism. The two inhibitors occupy the peptide substrate-binding site and interact with the protein through primarily hydrophobic contacts. Analyses of the inhibitorenzyme interactions provide valuable information for further improvement of antifungal inhibitors targeting NMT.
Organizational Affiliation:
State Key Laboratory of Molecular Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai, China.