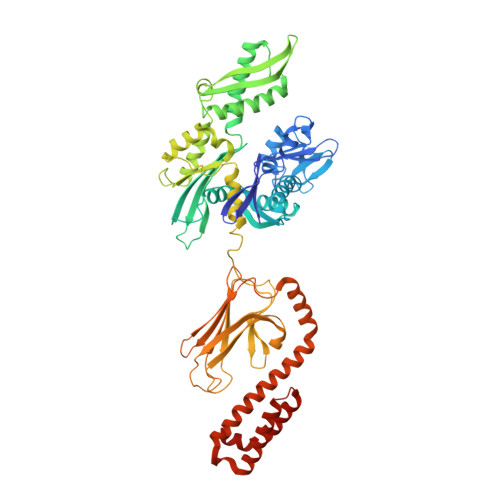
Solution conformation of wild-type E. coli Hsp70 (DnaK) chaperone complexed with ADP and substrate.
Bertelsen, E.B., Chang, L., Gestwicki, J.E., Zuiderweg, E.R.(2009) Proc Natl Acad Sci U S A 106: 8471-8476
- PubMed: 19439666
- DOI: https://doi.org/10.1073/pnas.0903503106
- Primary Citation of Related Structures:
2KHO - PubMed Abstract:
DnaK is the canonical Hsp70 molecular chaperone protein from Escherichia coli. Like other Hsp70s, DnaK comprises two main domains: a 44-kDa N-terminal nucleotide-binding domain (NBD) that contains ATPase activity, and a 25-kDa substrate-binding domain (SBD) that harbors the substrate-binding site. Here, we report an experimental structure for wild-type, full-length DnaK, complexed with the peptide NRLLLTG and with ADP. It was obtained in aqueous solution by using NMR residual dipolar coupling and spin labeling methods and is based on available crystal structures for the isolated NBD and SBD. By using dynamics methods, we determine that the NBD and SBD are loosely linked and can move in cones of +/-35 degrees with respect to each other. The linker region between the domains is a dynamic random coil. Nevertheless, an average structure can be defined. This structure places the SBD in close proximity of subdomain IA of the NBD and suggests that the SBD collides with the NBD at this area to establish allosteric communication.
Organizational Affiliation:
Department of Biological Chemistry, University of Michigan, Ann Arbor, MI 48109, USA.