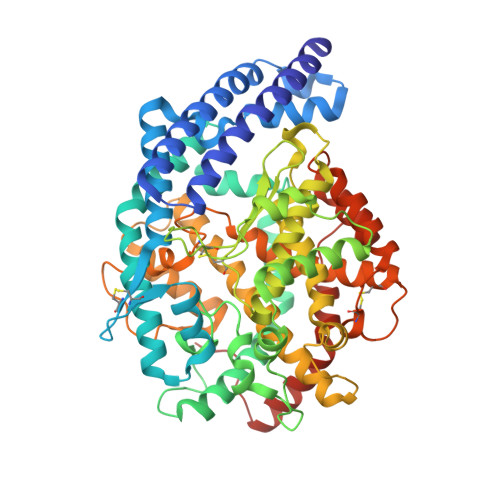
Structure of Testis Ace Glycosylation Mutants and Evidence for Conserved Domain Movement.
Watermeyer, J.M., Sewell, B.T., Schwager, S.L., Natesh, R., Corradi, H.R., Acharya, K.R., Sturrock, E.D.(2006) Biochemistry 45: 12654
- PubMed: 17042482
- DOI: https://doi.org/10.1021/bi061146z
- Primary Citation of Related Structures:
2IUL, 2IUX - PubMed Abstract:
Human angiotensin-converting enzyme is an important drug target for which little structural information has been available until recent years. The slow progress in obtaining a crystal structure was due to the problem of surface glycosylation, a difficulty that has thus far been overcome by the use of a glucosidase-1 inhibitor in the tissue culture medium. However, the prohibitive cost of these inhibitors and incomplete glucosidase inhibition makes alternative routes to minimizing the N-glycan heterogeneity desirable. Here, glycosylation in the testis isoform (tACE) has been reduced by Asn-Gln point mutations at N-glycosylation sites, and the crystal structures of mutants having two and four intact sites have been solved to 2.0 A and 2.8 A, respectively. Both mutants show close structural identity with the wild-type. A hinge mechanism is proposed for substrate entry into the active cleft, based on homology to human ACE2 at the levels of sequence and flexibility. This is supported by normal-mode analysis that reveals intrinsic flexibility about the active site of tACE. Subdomain II, containing bound chloride and zinc ions, is found to have greater stability than subdomain I in the structures of three ACE homologues. Crystallizable glycosylation mutants open up new possibilities for cocrystallization studies to aid the design of novel ACE inhibitors.
Organizational Affiliation:
Division of Medical Biochemistry, Institute of Infectious Disease and Molecular Medicine, University of Cape Town, South Africa.