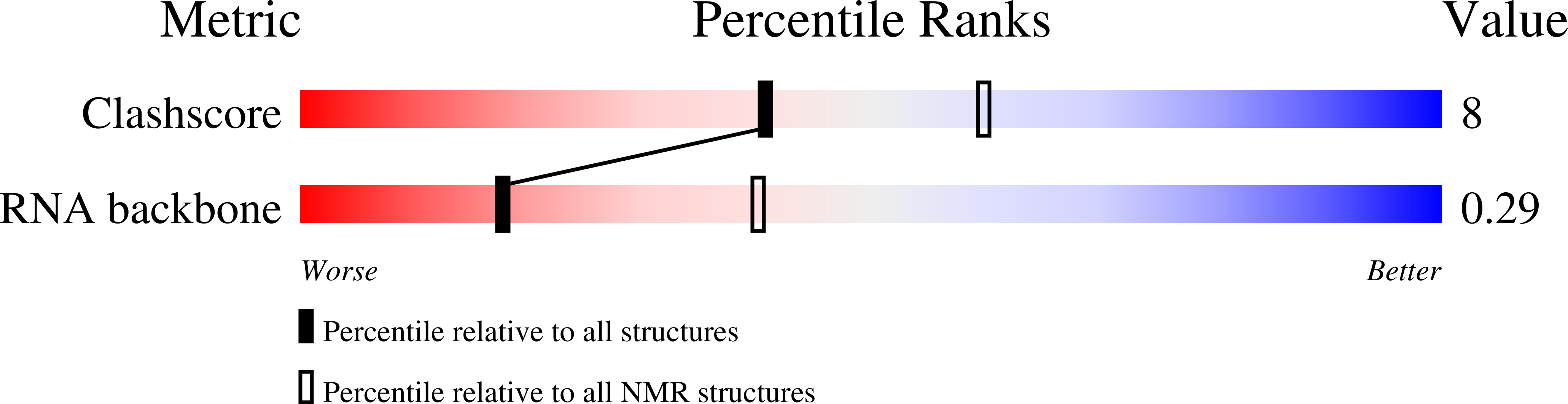
Breaking pseudo-twofold symmetry in the poliovirus 3'-UTR Y-stem by restoring Watson-Crick base pairs.
Zoll, J., Tessari, M., Van Kuppeveld, F.J.M., Melchers, W.J.G., Heus, H.A.(2007) RNA 13: 781-792
- PubMed: 17449731
- DOI: https://doi.org/10.1261/rna.375607
- Primary Citation of Related Structures:
2GRW, 2GV4 - PubMed Abstract:
The previously described NMR structure of a 5'-CU-3'/5'-UU-3' motif, which is highly conserved within the 3'-UTR Y-stem of poliovirus-like enteroviruses, revealed striking regularities of the local helix geometry, thus retaining the pseudo-twofold symmetry of the RNA helix. A mutant virus with both pyrimidine base pairs changed into Watson-Crick replicated as wild type, indicating the functional importance of this symmetry relation in viral RNA replication. Here we investigated the effect of changing only one of the two pyrimidine base pairs to Watson-Crick. We determined the NMR structures of two Y-stem variants: one containing the 5'-CU-3'/5'-AU-3' motif, which has been found in wild-type virus isolates as well, and the other containing a 5'-CU-3'/5'-UG-3' motif, which is not present in any enterovirus sequenced to date. Both structures show single pyrimidine mismatches with intercalated bases. In the 5'-CU-3'/5'-AU-3' motif a C-U Watson-Crick-type base pair is formed that retains the pseudo-twofold symmetry, while in the 5'-CU-3'/5'-UG-3' motif a single asymmetric U-U mismatch breaks the twofold symmetry. Surprisingly, for the nonnatural variant no effect of the single base-pair replacement was observed on polioviral RNA replication using an in vitro replicon assay.
Organizational Affiliation:
Institute for Molecules and Materials, Laboratory of Biophysical Chemistry, Radboud University Nijmegen, Toernooiveld 1, 6525 ED Nijmegen, The Netherlands.