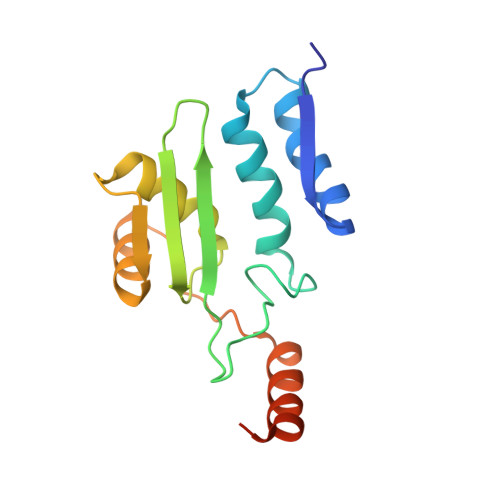
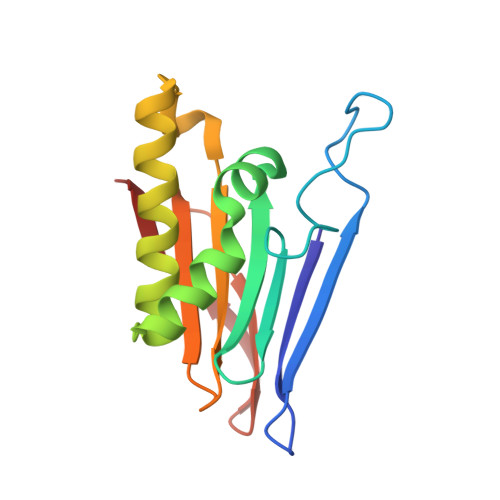
Crystallographic snapshot of a productive glycosylasparaginase-substrate complex.
Wang, Y., Guo, H.C.(2007) J Mol Biol 366: 82-92
- PubMed: 17157318
- DOI: https://doi.org/10.1016/j.jmb.2006.09.051
- Primary Citation of Related Structures:
2GL9 - PubMed Abstract:
Glycosylasparaginase (GA) plays an important role in asparagine-linked glycoprotein degradation. A deficiency in the activity of human GA leads to a lysosomal storage disease named aspartylglycosaminuria. GA belongs to a superfamily of N-terminal nucleophile hydrolases that autoproteolytically generate their mature enzymes from inactive single chain protein precursors. The side-chain of the newly exposed N-terminal residue then acts as a nucleophile during substrate hydrolysis. By taking advantage of mutant enzyme of Flavobacterium meningosepticum GA with reduced enzymatic activity, we have obtained a crystallographic snapshot of a productive complex with its substrate (NAcGlc-Asn), at 2.0 A resolution. This complex structure provided us an excellent model for the Michaelis complex to examine the specific contacts critical for substrate binding and catalysis. Substrate binding induces a conformational change near the active site of GA. To initiate catalysis, the side-chain of the N-terminal Thr152 is polarized by the free alpha-amino group on the same residue, mediated by the side-chain hydroxyl group of Thr170. Cleavage of the amide bond is then accomplished by a nucleophilic attack at the carbonyl carbon of the amide linkage in the substrate, leading to the formation of an acyl-enzyme intermediate through a negatively charged tetrahedral transition state.
Organizational Affiliation:
Department of Physiology and Biophysics, Boston University School of Medicine, 715 Albany Street, Boston, MA 02118-2526, USA.