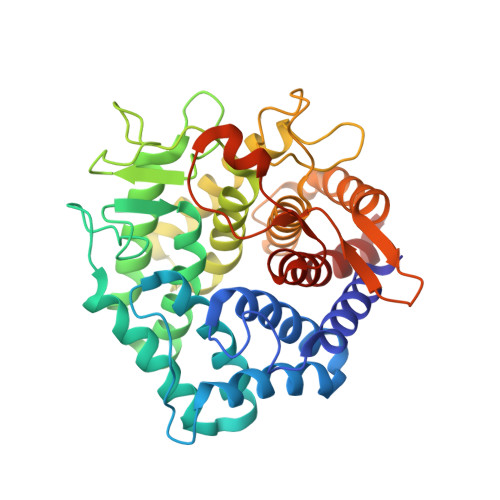
Structure of unsaturated rhamnogalacturonyl hydrolase complexed with substrate
Itoh, T., Ochiai, A., Mikami, B., Hashimoto, W., Murata, K.(2006) Biochem Biophys Res Commun 347: 1021-1029
- PubMed: 16870154
- DOI: https://doi.org/10.1016/j.bbrc.2006.07.034
- Primary Citation of Related Structures:
2GH4 - PubMed Abstract:
Bacillus subtilis strain 168 YteR has been identified as a novel enzyme "unsaturated rhamnogalacturonyl hydrolase" classified in glycoside hydrolase family 105. This enzyme acts specifically on unsaturated rhamnogalacturonan (RG) produced from plant cell wall RG type-I treated with RG lyases, releasing unsaturated galacturonic acid (DeltaGalA) from the substrate. The most likely candidate catalytic residue is Asp-143. Here, we show the structure of D143N in complex with unsaturated RG disaccharide (substrate) determined at 1.9A resolution by X-ray crystallography. This structural feature directly contributes to the postulation of the enzyme reaction mechanism. YteR triggers the hydration of vinyl ether group in DeltaGalA, but not of glycoside bond, by using Asp-143 as a general acid and base catalyst. Asp-143 donates proton to the double bond of DeltaGalA as an acid catalyst and also deprotonates a water molecule as a base catalyst. Deprotonated water molecule attacks the C5 atom of DeltaGalA.
Organizational Affiliation:
Division of Applied Life Sciences, Graduate School of Agriculture, Kyoto University, Gokasho, Uji, Kyoto 611-0011, Japan.