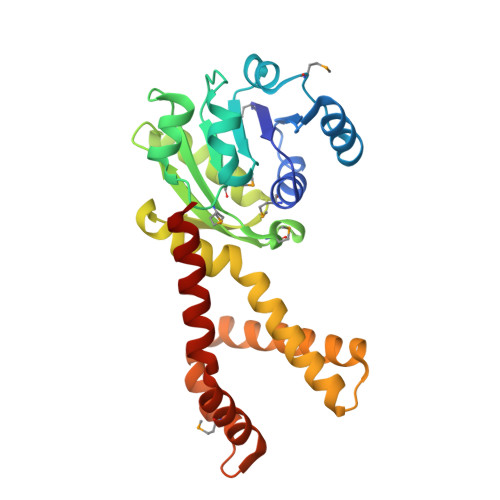
Crystal Structures of Delta(1)-Pyrroline-5-carboxylate Reductase from Human Pathogens Neisseria meningitides and Streptococcus pyogenes
Nocek, B., Chang, C., Li, H., Lezondra, L., Holzle, D., Collart, F., Joachimiak, A.(2005) J Mol Biol 354: 91-106
- PubMed: 16233902
- DOI: https://doi.org/10.1016/j.jmb.2005.08.036
- Primary Citation of Related Structures:
1YQG, 2AG8, 2AHR, 2AMF - PubMed Abstract:
L-proline is an amino acid that plays an important role in proteins uniquely contributing to protein folding, structure, and stability, and this amino acid serves as a sequence-recognition motif. Proline biosynthesis can occur via two pathways, one from glutamate and the other from arginine. In both pathways, the last step of biosynthesis, the conversion of delta1-pyrroline-5-carboxylate (P5C) to L-proline, is catalyzed by delta1-pyrroline-5-carboxylate reductase (P5CR) using NAD(P)H as a cofactor. We have determined the first crystal structure of P5CR from two human pathogens, Neisseria meningitides and Streptococcus pyogenes, at 2.0 angstroms and 2.15 angstroms resolution, respectively. The catalytic unit of P5CR is a dimer composed of two domains, but the biological unit seems to be species-specific. The N-terminal domain of P5CR is an alpha/beta/alpha sandwich, a Rossmann fold. The C-terminal dimerization domain is rich in alpha-helices and shows domain swapping. Comparison of the native structure of P5CR to structures complexed with L-proline and NADP+ in two quite different primary sequence backgrounds provides unique information about key functional features: the active site and the catalytic mechanism. The inhibitory L-proline has been observed in the crystal structure.
Organizational Affiliation:
Midwest Center for Structural Genomics and Structural Biology Center, Biosciences Division, Argonne National Laboratory, 9700 South Cass Avenue, Building 202, Argonne, IL 60439, USA.