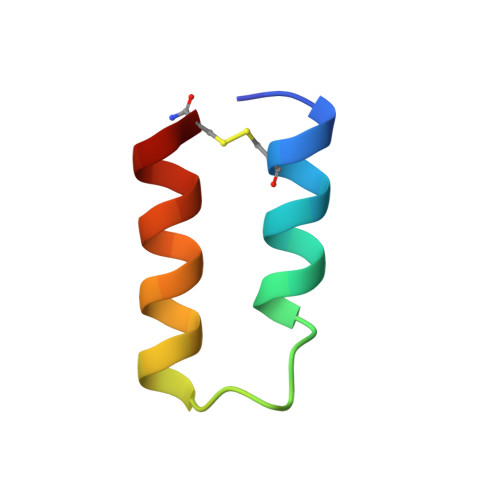
Structural mimicry of a native protein by a minimized binding domain.
Starovasnik, M.A., Braisted, A.C., Wells, J.A.(1997) Proc Natl Acad Sci U S A 94: 10080-10085
- PubMed: 9294166
- DOI: https://doi.org/10.1073/pnas.94.19.10080
- Primary Citation of Related Structures:
1ZDA, 1ZDB, 1ZDC, 1ZDD - PubMed Abstract:
The affinity between molecules depends both on the nature and presentation of the contacts. Here, we observe coupling of functional and structural elements when a protein binding domain is evolved to a smaller functional mimic. Previously, a 38-residue form of the 59-residue B-domain of protein A, termed Z38, was selected by phage display. Z38 contains 13 mutations and binds IgG only 10-fold weaker than the native B-domain. We present the solution structure of Z38 and show that it adopts a tertiary structure remarkably similar to that observed for the first two helices of B-domain in the B-domain/Fc complex [Deisenhofer, J. (1981) Biochemistry 20, 2361-2370], although it is significantly less stable. Based on this structure, we have improved on Z38 by designing a 34-residue disulfide-bonded variant (Z34C) that has dramatically enhanced stability and binds IgG with 9-fold higher affinity. The improved stability of Z34C led to NMR spectra with much greater chemical shift dispersion, resulting in a more precisely determined structure. Z34C, like Z38, has a structure virtually identical to the equivalent region from native protein A domains. The well-defined hydrophobic core of Z34C reveals key structural features that have evolved in this small, functional domain. Thus, the stabilized two-helix peptide, about half the size and having one-third of the remaining residues altered, accurately mimics both the structure and function of the native domain.
Organizational Affiliation:
Department of Protein Engineering, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA.