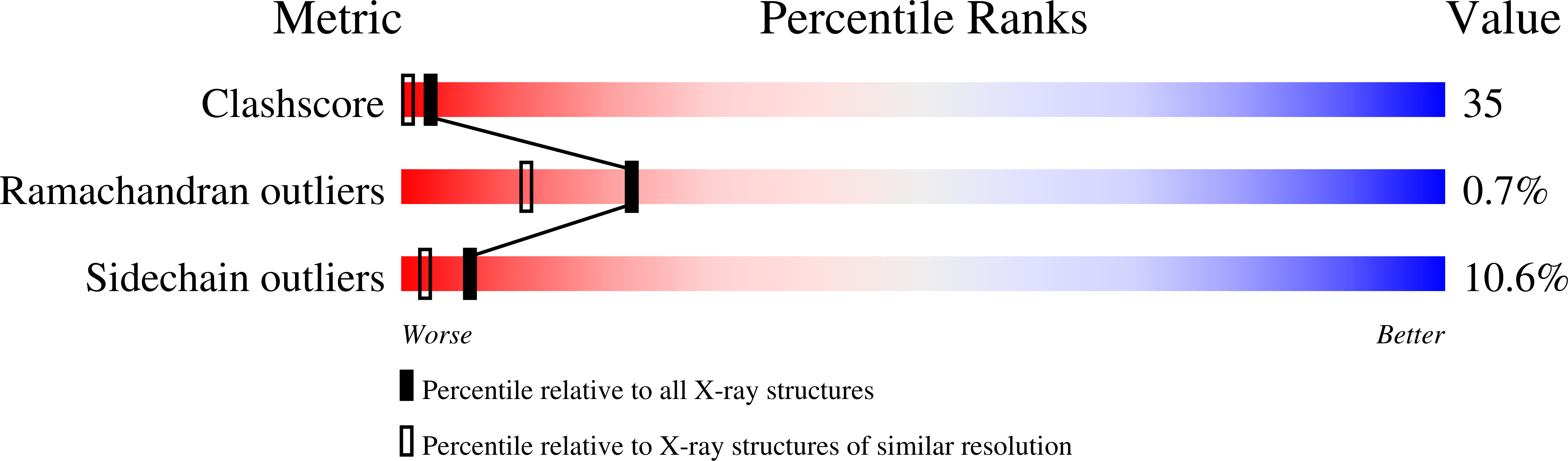High-resolution study of the three-dimensional structure of horse heart metmyoglobin.
Evans, S.V., Brayer, G.D.(1990) J Mol Biol 213: 885-897
- PubMed: 2359126
- DOI: https://doi.org/10.1016/S0022-2836(05)80270-0
- Primary Citation of Related Structures:
1YMB - PubMed Abstract:
The three-dimensional structure of horse heart metmyoglobin has been refined to a final R-factor of 15.5% for all observed data in the 6.0 to 1.9 A resolution range. The final model consists of 1242 non-hydrogen protein atoms, 154 water molecules and one sulfate ion. This structure has nearly ideal bonding and bond angle geometry. A Luzzati plot of the variation in R-factor with resolution yields an estimated mean co-ordinate error of 0.18 A. An extensive analysis of the pattern of hydrogen bonds formed in horse heart metmyoglobin has been completed. Over 80% of the polypeptide chain is involved in eight helical segments, of which seven are composed mainly of alpha-helical (3.6(13))-type hydrogen bonds; the remaining helix is composed entirely of 3(10) hydrogen bonds. Altogether, of 102 hydrogen bonds between main-chain atoms only six are not involved in helical structures, and four of these six occur within beta-turns. The majority of water molecules in horse heart metmyoglobin are found in solvent networks that range in size from two to 35 members. The size of water molecule networks can be rationalized on the basis of three factors: the number of hydrogen bonds to the protein surface, the presence of charged side-chain atoms, and the ability to bridge to neighboring molecules in the crystal lattice. Bridging water networks form the dominant intermolecular interactions. The backbone conformation of horse heart metmyoglobin is very similar to sperm whale metmyoglobin, with significant differences in secondary structure occurring only near residues 119 and 120, where residues 120 to 123 in sperm whale form a distorted type I reverse turn and the horse heart protein has a type II turn at residues 119 to 122. Nearly all of the hydrogen bonds between main-chain atoms (occurring mainly in helical regions) are common to both proteins, and more than half of the hydrogen bonds involving side-chain atoms observed in horse heart are also found in sperm whale metmyoglobin. Unlike sperm whale metmyoglobin, the heme iron atom in horse heart metmyoglobin is not significantly displaced from the plane of the heme group.
Organizational Affiliation:
Department of Biochemistry, University of British Columbia, Vancouver, Canada.