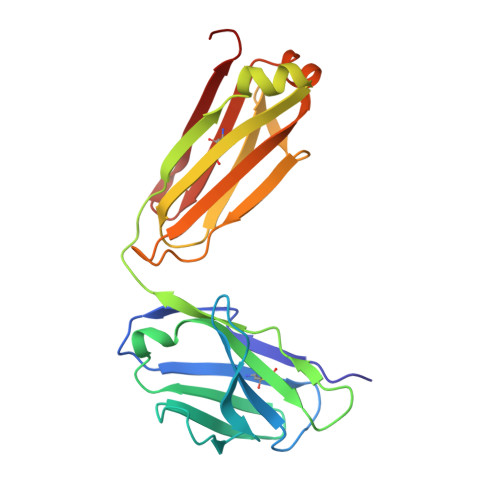
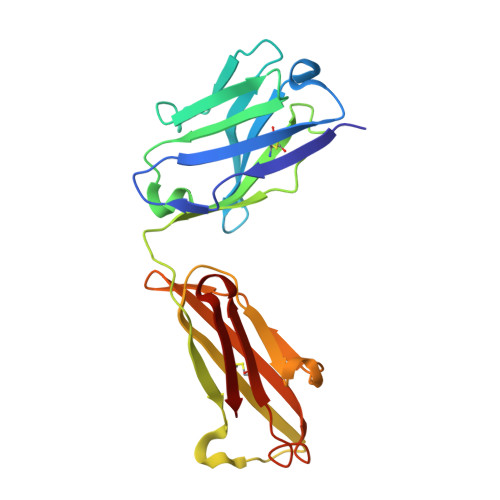
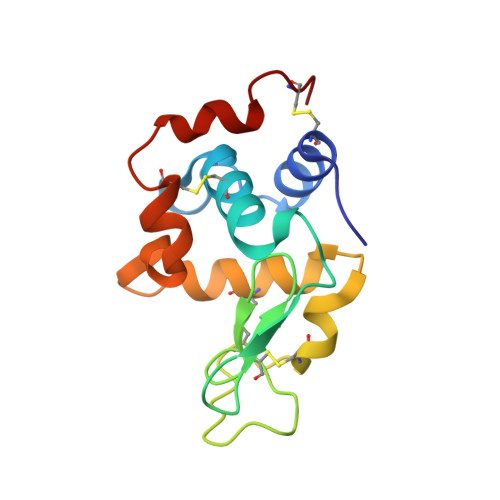
Magnitude of the hydrophobic effect at central versus peripheral sites in protein-protein interfaces
Li, Y., Huang, Y., Swaminathan, C.P., Smith-Gill, S.J., Mariuzza, R.A.(2005) Structure 13: 297-307
- PubMed: 15698573
- DOI: https://doi.org/10.1016/j.str.2004.12.012
- Primary Citation of Related Structures:
1XGP, 1XGQ, 1XGR, 1XGT, 1XGU - PubMed Abstract:
Hydrophobic interactions are essential for stabilizing protein-protein complexes, whose interfaces generally consist of a central cluster of hot spot residues surrounded by less important peripheral residues. According to the O-ring hypothesis, a condition for high affinity binding is solvent exclusion from interacting residues. This hypothesis predicts that the hydrophobicity at the center is significantly greater than at the periphery, which we estimated at 21 cal mol(-1) A(-2). To measure the hydrophobicity at the center, structures of an antigen-antibody complex where a buried phenylalanine was replaced by smaller hydrophobic residues were determined. By correlating structural changes with binding free energies, we estimate the hydrophobicity at this central site to be 46 cal mol(-1) A(-2), twice that at the periphery. This context dependence of the hydrophobic effect explains the clustering of hot spots at interface centers and has implications for hot spot prediction and the design of small molecule inhibitors.
Organizational Affiliation:
Center for Advanced Research in Biotechnology, W.M. Keck Laboratory for Structural Biology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, Maryland 20859, USA.