Characterization of a New Member of the Flavoprotein Disulfide Reductase Family of Enzymes from Mycobacterium tuberculosis
Argyrou, A., Vetting, M.W., Blanchard, J.S.(2004) J Biol Chem 279: 52694-52702
- PubMed: 15456792
- DOI: https://doi.org/10.1074/jbc.M410704200
- Primary Citation of Related Structures:
1XDI - PubMed Abstract:
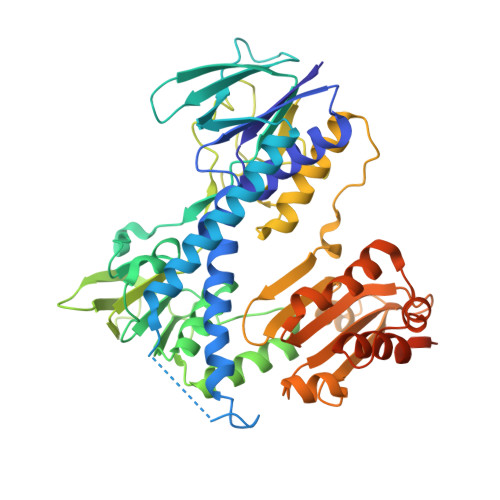
The lpdA (Rv3303c) gene from Mycobacterium tuberculosis encoding a new member of the flavoprotein disulfide reductases was expressed in Escherichia coli, and the recombinant LpdA protein was purified to homogeneity. LpdA is a homotetramer and co-purifies with one molecule of tightly but noncovalently bound FAD and NADP+ per monomer. Although annotated as a probable lipoamide dehydrogenase in M. tuberculosis, LpdA cannot catalyze reduction of lipoyl substrates, because it lacks one of two cysteine residues involved in dithiol-disulfide interchange with lipoyl substrates and a His-Glu pair involved in general acid catalysis. The crystal structure of LpdA was solved by multiple isomorphous replacement with anomalous scattering, which confirmed the absence of these catalytic residues from the active site. Although LpdA cannot catalyze reduction of disulfide-bonded substrates, it catalyzes the NAD(P)H-dependent reduction of alternative electron acceptors such as 2,6-dimethyl-1,4-benzoquinone and 5-hydroxy-1,4-naphthaquinone. Significant primary deuterium kinetic isotope effects were observed with [4S-2H]NADH establishing that the enzyme promotes transfer of the C4-proS hydride of NADH. The absence of an isotope effect with [4S-2H]NADPH, the low Km value of 0.5 microm for NADPH, and the potent inhibition of the NADH-dependent reduction of 2,6-dimethyl-1,4-benzoquinone by NADP+ (Ki approximately 6 nm) and 2'-phospho-ADP-ribose (Ki approximately 800 nm), demonstrate the high affinity of LpdA for 2'-phosphorylated nucleotides and that the physiological substrate/product pair is NADPH/NADP+ rather than NADH/NAD+. Modeling of NADP+ in the active site revealed that LpdA achieves the high specificity for NADP+ through interactions involving the 2'-phosphate of NADP+ and amino acid residues that are different from those in glutathione reductase.
Organizational Affiliation:
Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York 10461, USA. aargyrou@medusa.bioc.aecom.yu.edu