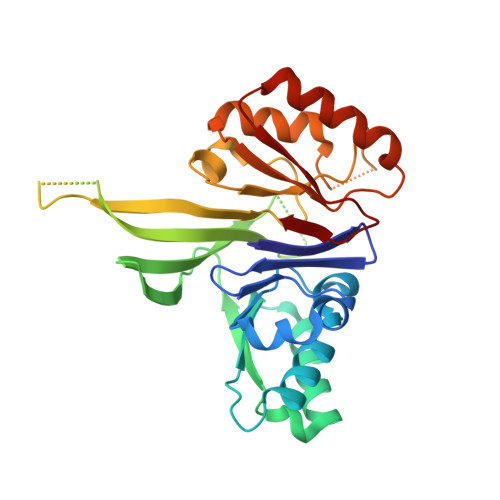
Crystal Structure of the tRNA 3' Processing Endoribonuclease tRNase Z from Thermotoga maritima
Ishii, R., Minagawa, A., Takaku, H., Takagi, M., Nashimoto, M., Yokoyama, S.(2005) J Biol Chem 280: 14138-14144
- PubMed: 15701599
- DOI: https://doi.org/10.1074/jbc.M500355200
- Primary Citation of Related Structures:
1WW1 - PubMed Abstract:
The maturation of the tRNA 3' end is catalyzed by a tRNA 3' processing endoribonuclease named tRNase Z (RNase Z or 3'-tRNase) in eukaryotes, Archaea, and some bacteria. The tRNase Z generally cuts the 3' extra sequence from the precursor tRNA after the discriminator nucleotide. In contrast, Thermotoga maritima tRNase Z cleaves the precursor tRNA precisely after the CCA sequence. In this study, we determined the crystal structure of T. maritima tRNase Z at 2.6-A resolution. The tRNase Z has a four-layer alphabeta/betaalpha sandwich fold, which is classified as a metallo-beta-lactamase fold, and forms a dimer. The active site is located at one edge of the beta-sandwich and is composed of conserved motifs. Based on the structure, we constructed a docking model with the tRNAs that suggests how tRNase Z may recognize the substrate tRNAs.
Organizational Affiliation:
Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.