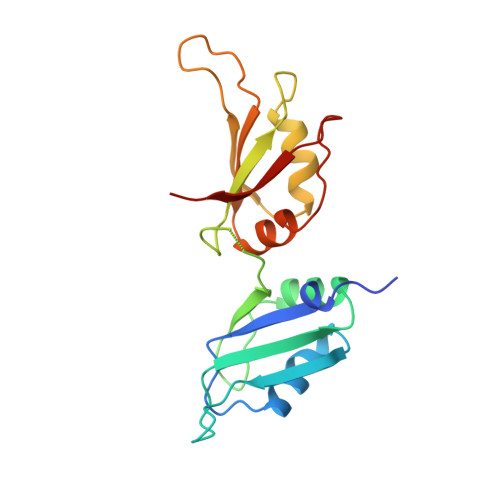
Crystal structure of human UP1, the domain of hnRNP A1 that contains two RNA-recognition motifs.
Xu, R.M., Jokhan, L., Cheng, X., Mayeda, A., Krainer, A.R.(1997) Structure 5: 559-570
- PubMed: 9115444
- DOI: https://doi.org/10.1016/s0969-2126(97)00211-6
- Primary Citation of Related Structures:
1UP1 - PubMed Abstract:
Heterogeneous nuclear ribonucleoprotein (hnRNP) A1 is one of the most abundant core proteins of hnRNP complexes in metazoan nuclei. It behaves as a global regulator of alternative pre-mRNA splicing by antagonizing the activities of several serine/arginine-rich splicing factors (SR proteins), resulting in the activation of distal alternative 5' splice sites and skipping of optional exons. Purified hnRNP A1 has nucleic acid annealing activity. The protein also shuttles continuously between the nucleus and the cytoplasm, a process mediated by signals within its C-terminal glycine-rich domain. The N-terminal region of human hnRNP A1, termed unwinding protein 1 (UP1), contains two RNA-recognition motifs (RRMs), RRM1 and RRM2. Understanding the structural elements by which hnRNP A1 interacts with RNA will have broad implications for studies of RNA processing. The crystal structure of UP1 has been determined to 1.9 A resolution. Each RRM independently adopts the characteristic RRM fold, consisting of a four-stranded antiparallel beta-pleated sheet and two alpha helices packed on one side of the beta sheet. The two RRMs are antiparallel and held in close contact, mainly by two Arg-Asp ion pairs. As a result, the two four-stranded beta sheets are brought together to form an extended RNA-binding surface. A segment of the linker connecting the two RRMs is flexible in the absence of bound RNA, but the general location of the linker suggests that it can make direct contacts with RNA. Comparison with other RRM structures indicates that a short 310 helix, found immediately N-terminal to the first beta strand in RRM1, may interact with RNA directly. The RRM is one of the most common and best characterized RNA-binding motifs. In certain cases, one RRM is sufficient for sequence-specific and high affinity RNA binding; but in other cases, synergy between several RRMs within a single protein is required. This study shows how two RRMs are organized in a single polypeptide. The two independently folded RRMs in UP1 are held together in a fixed geometry, enabling the two RRMs to function as a single entity in binding RNA, and so explaining the synergy between the RRMs. The UP1 structure also suggests that residues which lie outside of the RRMs can make potentially important interactions with RNA.
Organizational Affiliation:
WM Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, P.O. Box 100, Cold Spring Harbor, New York, 11724, USA. xur@cshl.org