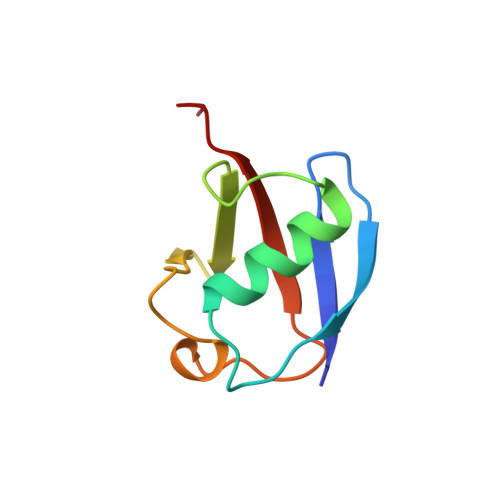
Solution structure and dynamics of a designed hydrophobic core variant of ubiquitin.
Johnson, E.C., Lazar, G.A., Desjarlais, J.R., Handel, T.M.(1999) Structure 7: 967-976
- PubMed: 10467150
- DOI: https://doi.org/10.1016/s0969-2126(99)80123-3
- Primary Citation of Related Structures:
1UD7 - PubMed Abstract:
The recent merger of computation and protein design has resulted in a burst of success in the generation of novel proteins with native-like properties. A critical component of this coupling between theory and experiment is a detailed analysis of the structures and stabilities of designed proteins to assess and improve the accuracy of design algorithms. Here we report the solution structure of a hydrophobic core variant of ubiquitin, referred to as 1D7, which was designed with the core-repacking algorithm ROC. As a measure of conformational specificity, we also present amide exchange protection factors and backbone and sidechain dynamics. The results indicate that 1D7 is similar to wild-type (WT) ubiquitin in backbone structure and degree of conformational specificity. We also observe a good correlation between experimentally determined sidechain structures and those predicted by ROC. However, evaluation of the core sidechain conformations indicates that, in general, 1D7 has more sidechains in less statistically favorable conformations than WT. Our results provide an explanation for the lower stability of 1D7 compared to WT, and suggest modifications to design algorithms that may improve the accuracy with which structure and stability are predicted. The results also demonstrate that core packing can affect conformational flexibility in subtle ways that are likely to be important for the design of function and protein-ligand interactions.
Organizational Affiliation:
Department of Physics, University of California, Berkeley 94720, USA.