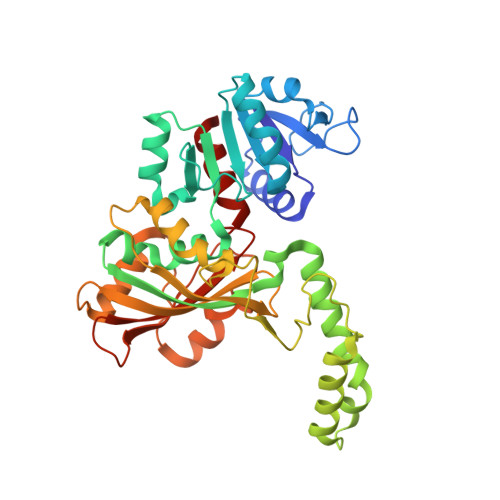
High-resolution Structures Reveal Details of Domain Closure and "Half-of-sites-reactivity" in Escherichia coli Aspartate beta-Semialdehyde Dehydrogenase.
Nichols, C.E., Dhaliwal, B., Lockyer, M., Hawkins, A.R., Stammers, D.K.(2004) J Mol Biol 341: 797-806
- PubMed: 15288787
- DOI: https://doi.org/10.1016/j.jmb.2004.05.073
- Primary Citation of Related Structures:
1T4B, 1T4D - PubMed Abstract:
Two high-resolution structures have been determined for Eschericia coli aspartate beta-semialdehyde dehydrogenase (ecASADH), an enzyme of the aspartate biosynthetic pathway, which is a potential target for novel antimicrobial drugs. Both ASADH structures were of the open form and were refined to 1.95 A and 1.6 A resolution, allowing a more detailed comparison with the closed form of the enzyme than previously possible. A more complex scheme for domain closure is apparent with the subunit being split into two further sub-domains with relative motions about three hinge axes. Analysis of hinge data and torsion-angle difference plots is combined to allow the proposal of a detailed structural mechanism for ecASADH domain closure. Additionally, asymmetric distortions of individual subunits are identified, which form the basis for the previously reported "half-of-the-sites reactivity" (HOSR). A putative explanation of this arrangement is also presented, suggesting the HOSR system may provide a means for ecASADH to offset the energy required to remobilise flexible loops at the end of the reaction cycle, and hence avoid falling into an energy minimum.
Organizational Affiliation:
Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford, OX3 7BN, UK.