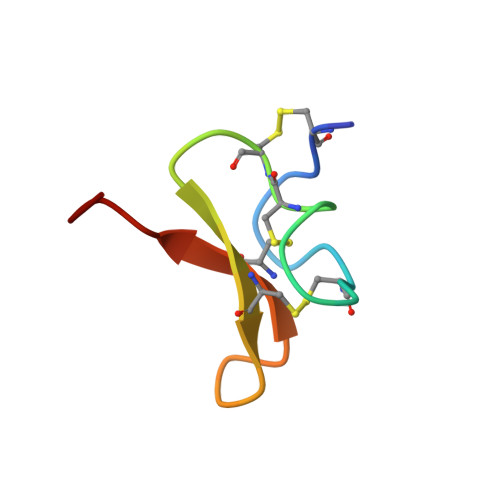
Solution structure and lipid membrane partitioning of VSTx1, an inhibitor of the KvAP potassium channel.
Jung, H.J., Lee, J.Y., Kim, S.H., Eu, Y.J., Shin, S.Y., Milescu, M., Swartz, K.J., Kim, J.I.(2005) Biochemistry 44: 6015-6023
- PubMed: 15835890
- DOI: https://doi.org/10.1021/bi0477034
- Primary Citation of Related Structures:
1S6X - PubMed Abstract:
VSTx1 is a voltage sensor toxin from the spider Grammostola spatulata that inhibits KvAP, an archeabacterial voltage-activated K(+) channel whose X-ray structure has been reported. Although the receptor for VSTx1 and the mechanism of inhibition are unknown, the sequence of the toxin is related to hanatoxin (HaTx) and SGTx, two toxins that inhibit eukaryotic voltage-activated K(+) channels by binding to voltage sensors. VSTx1 has been recently shown to interact equally well with lipid membranes that contain zwitterionic or acidic phospholipids, and it has been proposed that the toxin receptor is located within a region of the channel that is submerged in the membrane. As a first step toward understanding the inhibitory mechanism of VSTx1, we determined the three-dimensional solution structure of the toxin using NMR. Although the structure of VSTx1 is similar to HaTx and SGTx in terms of molecular fold and amphipathic character, the detailed positions of hydrophobic and surrounding charged residues in VSTx1 are very different than what is seen in the other toxins. The amphipathic character of VSTx1, notably the close apposition of basic and hydrophobic residues on one face of the toxin, raises the possibility that the toxin interacts with interfacial regions of the membrane. We reinvestigated the partitioning of VSTx1 into lipid membranes and find that VSTx1 partitioning requires negatively charged phospholipids. Intrinsic tryptophan fluorescence and acrylamide quenching experiments suggest that tryptophan residues on the hydrophobic surface of VSTx1 have a diminished exposure to water when the toxin interacts with membranes. The present results suggest that if membrane partitioning is involved in the mechanism by which VSTx1 inhibits voltage-activated K(+) channels, then binding of the toxin to the channel would likely occur at the interface between the polar headgroups and the hydrophobic phase of the membrane.
Organizational Affiliation:
Department of Life Science, Gwangju Institute of Science and Technology, Gwangju 500-712, Korea.