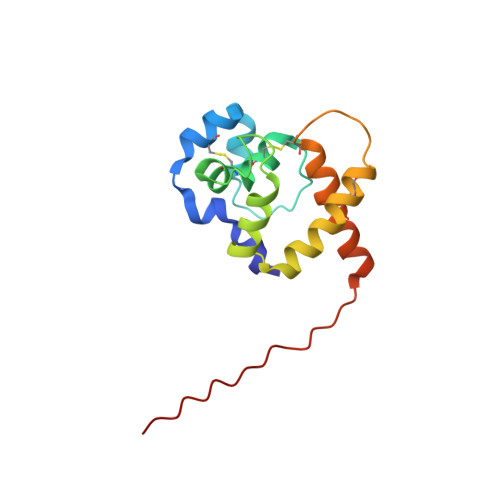
The Solution NMR Structure of Antheraea polyphemus PBP Provides New Insight into Pheromone Recognition by Pheromone-binding Proteins
Mohanty, S., Zubkov, S., Gronenborn, A.M.(2004) J Mol Biol 337: 443-451
- PubMed: 15003458
- DOI: https://doi.org/10.1016/j.jmb.2004.01.009
- Primary Citation of Related Structures:
1QWV - PubMed Abstract:
Pheromone-binding proteins (PBPs) located in the antennae of male moth species play an important role in olfaction. They are carrier proteins, believed to transport volatile hydrophobic pheromone molecules across the aqueous sensillar lymph to the membrane-bound G protein-coupled olfactory receptor proteins. The roles of PBPs in molecular recognition and the mechanisms of pheromone binding and release are poorly understood. Here, we report the NMR structure of a PBP from the giant silk moth Antheraea polyphemus. This is the first structure of a PBP with specific acetate-binding function in vivo. The protein consists of nine alpha-helices: alpha1a (residues 2-5), alpha1b (8-12), alpha1c (16-23), alpha2 (27-34), alpha3a (46-52), alpha3b (54-59), alpha4 (70-79), alpha5 (84-100) and alpha6 (107-125), held together by three disulfide bridges: 19-54, 50-108 and 97-117. A large hydrophobic cavity is located inside the protein, lined with side-chains from all nine helices. The acetate-binding site is located at the narrow end of the cavity formed by the helices alpha3b and alpha4. The pheromone can enter this cavity through an opening between the helix alpha1a, the C-terminal end of the helix alpha6, and the loop between alpha2 and alpha3a. We suggest that Trp37 may play an important role in the initial interaction with the ligand. Our analysis also shows that Asn53 plays the key role in recognition of acetate pheromones specifically, while Phe12, Phe36, Trp37, Phe76, and Phe118 are responsible for non-specific binding, and Leu8 and Ser9 may play a role in ligand chain length recognition.
Organizational Affiliation:
Department of Biochemistry and Cell Biology, State University of New York at Stony Brook, Stony Brook, NY 11794-5215, USA. smita.mohanty@sunysb.edu