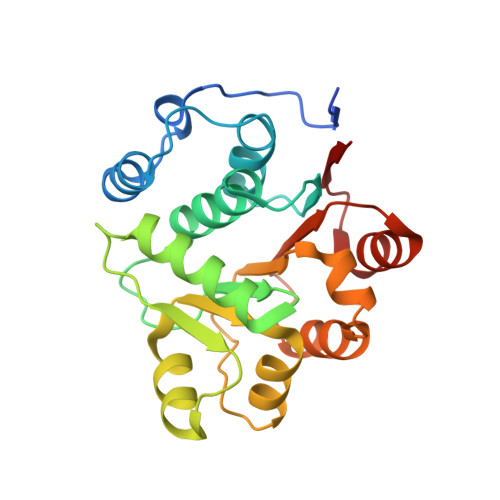
Crystallographic structure of the amino terminal domain of yeast initiation factor 4A, a representative DEAD-box RNA helicase
Johnson, E.R., McKay, D.B.(1999) RNA 5: 1526-1534
- PubMed: 10606264
- DOI: https://doi.org/10.1017/s1355838299991410
- Primary Citation of Related Structures:
1QVA - PubMed Abstract:
The eukaryotic translation initiation factor 4A (elF4A) is a representative of the DEAD-box RNA helicase protein family. We have solved the crystallographic structure of the amino-terminal domain (residues 1-223) of yeast elF4A. The domain is built around a core scaffold, a parallel alpha-beta motif with five beta strands, that is found in other RNA and DNA helicases, as well as in the RecA protein. The amino acid sequence motifs that are conserved within the helicase family are localized to the beta strand-->alpha helix junctions within the core. The core of the amino terminal domain of elF4A is amplified with additional structural elements that differ from those of other helicases. The phosphate binding loop (the Walker A motif) is in an unusual closed conformation. The crystallographic structure reveals specific interactions between amino acid residues of the phosphate binding loop, the DEAD motif, and the SAT motif, whose alteration is known to impair coupling between the ATPase cycle and the RNA unwinding activity of elF4A.
Organizational Affiliation:
Department of Structural Biology, Stanford University School of Medicine, California 94305-5400, USA.