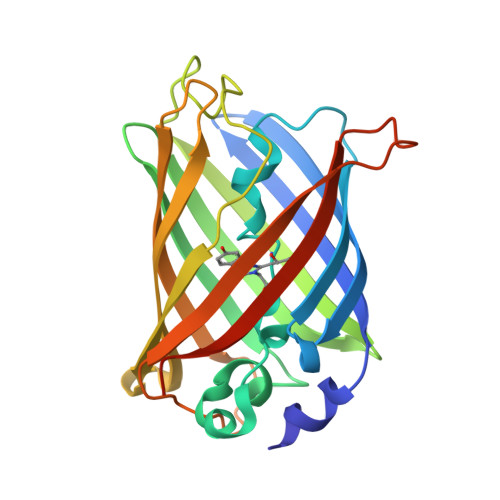
Local complexity of amino acid interactions in a protein core.
Jain, R.K., Ranganathan, R.(2004) Proc Natl Acad Sci U S A 101: 111-116
- PubMed: 14684834
- DOI: https://doi.org/10.1073/pnas.2534352100
- Primary Citation of Related Structures:
1Q4A, 1Q4B, 1Q4C, 1Q4D, 1Q4E, 1Q73 - PubMed Abstract:
Atomic resolution structures of proteins indicate that the core is typically well packed, suggesting a densely connected network of interactions between amino acid residues. The combinatorial complexity of energetic interactions in such a network could be enormous, a problem that limits our ability to relate structure and function. Here, we report a case study of the complexity of amino acid interactions in a localized region within the core of the GFP, a particularly stable and tightly packed molecule. Mutations at three sites within the chromophore-binding pocket display an overlapping pattern of conformational change and are thermodynamically coupled, seemingly consistent with the dense network model. However, crystallographic and energetic analyses of coupling between mutations paint a different picture; pairs of mutations couple through independent "hotspots" in the region of structural overlap. The data indicate that, even in highly stable proteins, the core contains sufficient plasticity in packing to uncouple high-order energetic interactions of residues, a property that is likely general in proteins.
Organizational Affiliation:
Howard Hughes Medical Institute and Department of Pharmacology, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, TX 75390-9050, USA.