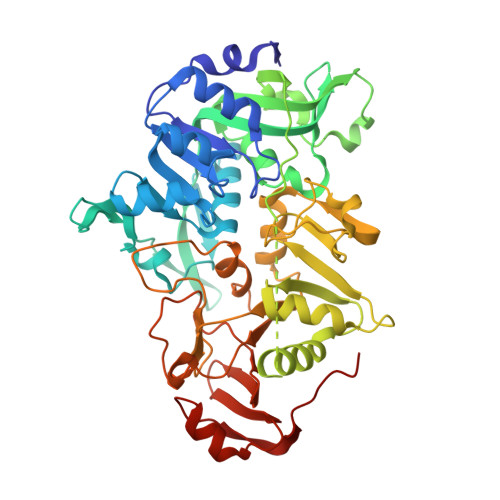
Structure of the CoA transferase from pig heart to 1.7 A resolution.
Coros, A.M., Swenson, L., Wolodko, W.T., Fraser, M.E.(2004) Acta Crystallogr D Biol Crystallogr 60: 1717-1725
- PubMed: 15388917
- DOI: https://doi.org/10.1107/S0907444904017974
- Primary Citation of Related Structures:
1OOY, 1OOZ, 1OPE - PubMed Abstract:
Succinyl-CoA:3-ketoacid CoA transferase (SCOT; EC 2.8.3.5) activates the acetoacetate in ketone bodies by transferring the CoA group from succinyl-CoA to acetoacetate to produce acetoacetyl-CoA and succinate. In the reaction, a glutamate residue at the active site of the enzyme forms a thioester bond with CoA and in this form the enzyme is subject to autolytic fragmentation. The crystal structure of pig heart SCOT has been solved and refined to 1.7 A resolution in a new crystal form. The structure shows the active-site glutamate residue in a conformation poised for autolytic fragmentation, with its side chain accepting one hydrogen bond from Asn281 and another from its own amide N atom. However, the conformation of this glutamate side chain would have to change for the residues that are conserved in the CoA transferases (Gln99, Gly386 and Ala387) to participate in stabilizing the tetrahedral transition states of the catalytic mechanism. The structures of a deletion mutant in two different crystal forms were also solved.
Organizational Affiliation:
Department of Biological Sciences, University of Calgary, Calgary, AB T2N 1N4, Canada.