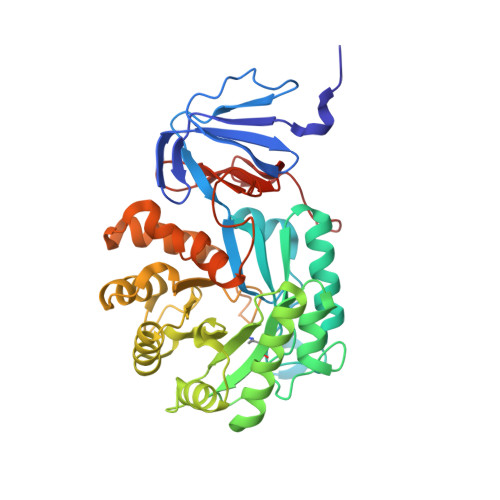
High Resolution X-ray Structure of Isoaspartyl Dipeptidase from Escherichia coli
Thoden, J.B., Marti-Arbona, R., Raushel, F.M., Holden, H.M.(2003) Biochemistry 42: 4874-4882
- PubMed: 12718528
- DOI: https://doi.org/10.1021/bi034233p
- Primary Citation of Related Structures:
1ONW, 1ONX - PubMed Abstract:
Isoaspartyl dipeptidase from Escherichia coli functions in protein degradation by catalyzing the hydrolysis of beta-L-isoaspartyl linkages in dipeptides. The best substrate for the enzyme reported thus far is iso-Asp-Leu. Here we report the X-ray analysis of the enzyme in its resting state and complexed with aspartate to 1.65 and 2.1 A resolution, respectively. The quaternary structure of the enzyme is octameric and can be aptly described as a tetramer of dimers. Each subunit folds into two distinct domains: the N-terminal region containing eight strands of mixed beta-sheet and the C-terminal motif that is dominated by a (beta,alpha)(8)-barrel. A binuclear zinc center is located in each subunit at the C-terminal end of the (beta,alpha)(8)-barrel. Ligands to the binuclear metal center include His 68, His 70, His 201, His 230, and Asp 285. The two zincs are bridged by a carboxylated lysine residue (Lys 162) and a solvent molecule, most likely a hydroxide ion. The product of the reaction, aspartate, binds to the enzyme by displacing the bridging solvent with its side chain functional group. From this investigation it is proposed that the reaction mechanism of the enzyme proceeds through a tetrahedral intermediate and that the bridging solvent attacks the re face of the carbonyl carbon of the scissile peptide bond. This structural analysis confirms the placement of isoaspartyl dipeptidase into the urease-related amidohydrolase superfamily.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.