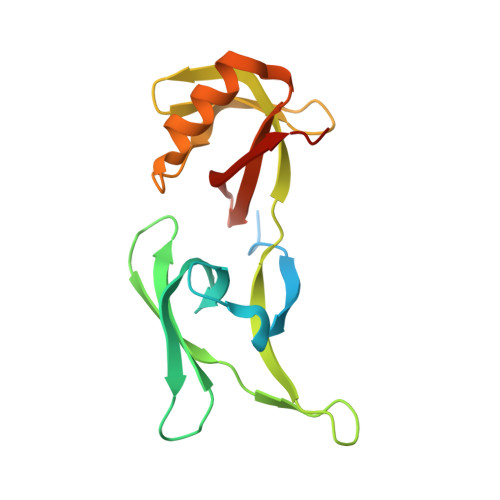
A HEX-1 crystal lattice required for Woronin body function in Neurospora crassa
Yuan, P., Jedd, G., Kumaran, D., Swaminathan, S., Shio, H., Hewitt, D., Chua, N.-H., Swaminathan, K.(2003) Nat Struct Biol 10: 264-270
- PubMed: 12640443
- DOI: https://doi.org/10.1038/nsb910
- Primary Citation of Related Structures:
1KHI - PubMed Abstract:
The Woronin body is a dense-core vesicle specific to filamentous ascomycetes (Euascomycetes), where it functions to seal the septal pore in response to cellular damage. The HEX-1 protein self-assembles to form this solid core of the vesicle. Here, we solve the crystal structure of HEX-1 at 1.8 A, which provides the structural basis of its self-assembly. The structure reveals the existence of three intermolecular interfaces that promote the formation of a three-dimensional protein lattice. Consistent with these data, self-assembly is disrupted by mutations in intermolecular contact residues and expression of an assembly-defective HEX-1 mutant results in the production of aberrant Woronin bodies, which possess a soluble noncrystalline core. This mutant also fails to complement a hex-1 deletion in Neurospora crassa, demonstrating that the HEX-1 protein lattice is required for Woronin body function. Although both the sequence and the tertiary structure of HEX-1 are similar to those of eukaryotic initiation factor 5A (eIF-5A), the amino acids required for HEX-1 self-assembly and peroxisomal targeting are absent in eIF-5A. Thus, we propose that a new function has evolved following duplication of an ancestral eIF-5A gene and that this may define an important step in fungal evolution.
Organizational Affiliation:
Institute of Molecular and Cell Biology, 30 Medical Drive, The National University of Singapore, Singapore, 117609.