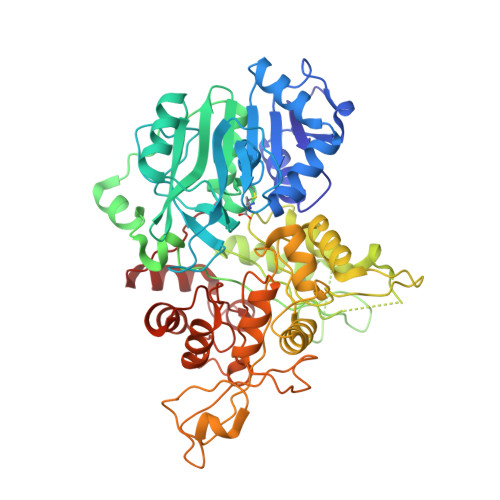
Crystal structure of imidazole glycerol phosphate synthase: a tunnel through a (beta/alpha)8 barrel joins two active sites.
Chaudhuri, B.N., Lange, S.C., Myers, R.S., Chittur, S.V., Davisson, V.J., Smith, J.L.(2001) Structure 9: 987-997
- PubMed: 11591353
- Primary Citation of Related Structures:
1JVN - PubMed Abstract:
Imidazole glycerol phosphate synthase catalyzes a two-step reaction of histidine biosynthesis at the bifurcation point with the purine de novo pathway. The enzyme is a new example of intermediate channeling by glutamine amidotransferases in which ammonia generated by hydrolysis of glutamine is channeled to a second active site where it acts as a nucleophile. In this case, ammonia reacts in a cyclase domain to produce imidazole glycerol phosphate and an intermediate of purine biosynthesis. The enzyme is also a potential target for drug and herbicide development since the histidine pathway does not occur in mammals. The 2.1 A crystal structure of imidazole glycerol phosphate synthase from yeast reveals extensive interaction of the glutaminase and cyclase catalytic domains. At the domain interface, the glutaminase active site points into the bottom of the (beta/alpha)(8) barrel of the cyclase domain. An ammonia tunnel through the (beta/alpha)(8) barrel connects the glutaminase docking site at the bottom to the cyclase active site at the top. A conserved "gate" of four charged residues controls access to the tunnel. This is the first structure in which all the components of the ubiquitous (beta/alpha)(8) barrel fold, top, bottom, and interior, take part in enzymatic function. Intimate contacts between the barrel domain and the glutaminase active site appear to be poised for crosstalk between catalytic centers in response to substrate binding at the cyclase active site. The structure provides a number of potential sites for inhibitor development in the active sites and in a conserved interdomain cavity.
Organizational Affiliation:
Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.