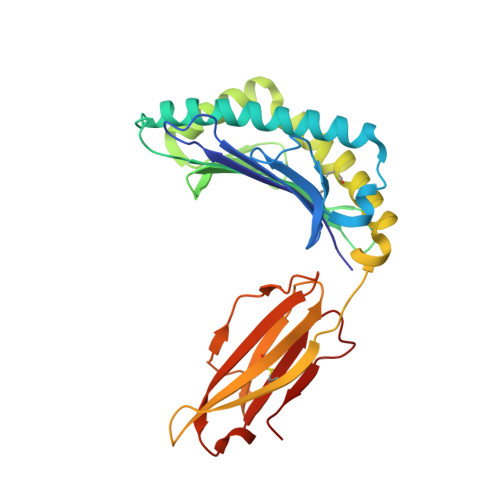
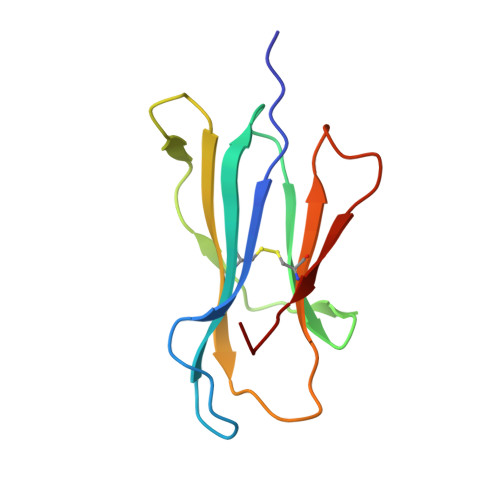
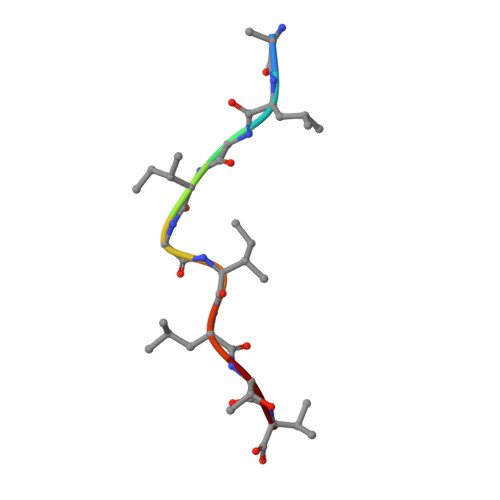
Crystal structures of two closely related but antigenically distinct HLA-A2/melanocyte-melanoma tumor-antigen peptide complexes.
Sliz, P., Michielin, O., Cerottini, J.C., Luescher, I., Romero, P., Karplus, M., Wiley, D.C.(2001) J Immunol 167: 3276-3284
- PubMed: 11544315
- DOI: https://doi.org/10.4049/jimmunol.167.6.3276
- Primary Citation of Related Structures:
1JF1, 1JHT - PubMed Abstract:
We have determined high-resolution crystal structures of the complexes of HLA-A2 molecules with two modified immunodominant peptides from the melanoma tumor-associated protein Melan-A/Melanoma Ag recognized by T cells-1. The two peptides, a decamer and nonamer with overlapping sequences (ELAGIGILTV and ALGIGILTV), are modified in the second residue to increase their affinity for HLA-A2. The modified decamer is more immunogenic than the natural peptide and a candidate for peptide-based melanoma immunotherapy. The crystal structures at 1.8 and 2.15 A resolution define the differences in binding modes of the modified peptides, including different clusters of water molecules that appear to stabilize the peptide-HLA interaction. The structures suggest both how the wild-type peptides would bind and how three categories of cytotoxic T lymphocytes with differing fine specificity might recognize the two peptides.
Organizational Affiliation:
Department of Molecular and Cellular Biology and Howard Hughes Medical Institute, Harvard University, Cambridge, MA 02138, USA.