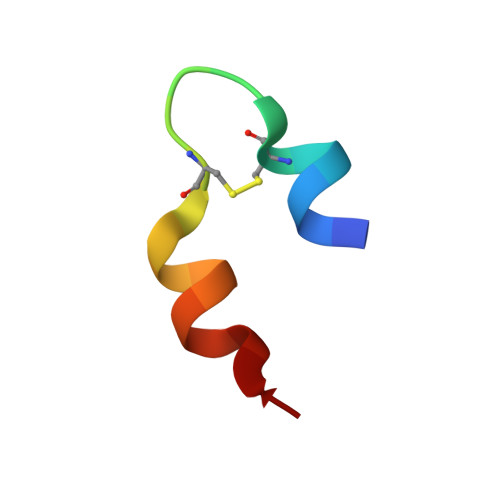
Flexibility and bioactivity of insulin: an NMR investigation of the solution structure and folding of an unusually flexible human insulin mutant with increased biological activity.
Keller, D., Clausen, R., Josefsen, K., Led, J.J.(2001) Biochemistry 40: 10732-10740
- PubMed: 11524020
- DOI: https://doi.org/10.1021/bi0108150
- Primary Citation of Related Structures:
1JCO - PubMed Abstract:
The structure and folding of a novel human insulin mutant, [Thr(B27) --> Pro, Pro(B28) --> Thr]insulin (PT insulin), in aqueous solution and in mixtures of water and 2,2,2-trifluoroethanol (TFE) have been studied by NMR spectroscopy. It was found that PT insulin has a highly flexible structure in pure water and is present in at least two different conformations, although with an overall tertiary structure similar to that of native insulin. Furthermore, the native helical structures are poorly defined. Surprisingly, the mutant has a biological activity about 50% higher than native insulin. In contrast, in TFE/water solution the mutant reveals a propensity of forming a well-defined structure at the secondary structure level, similar to monomeric native insulin. Thus, as shown by a detailed determination of the structure from 208 distance restraints and 52 torsion angle restraints by distance geometry, simulated annealing, and restrained energy minimization, the native insulin helices (A2-A7, A13-A19, and B10-B19) as well as the beta-turn (B20-B23) are formed in 35% TFE. However, the amount of tertiary structure is decreased significantly in TFE/water solution. The obtained results suggest that only an overall tertiary fold, as observed for PT insulin in pure water, is necessary for expressing the biological activity of insulin, as long as the molecule is flexible and retains the propensity to form the secondary structure required for its receptor binding. In contrast, a compact secondary structure, as found for native insulin in solution, is unnecessary for the biological activity. A model for the receptor binding of insulin is suggested that relates the increased bioactivity to the enhanced flexibility of the mutant.
Organizational Affiliation:
Department of Chemistry, University of Copenhagen, The H. C. Ørsted Institute, Universitetsparken 5, DK-2100 Copenhagen Ø, Denmark.