Crystal structures of two I-Ad-peptide complexes reveal that high affinity can be achieved without large anchor residues.
Scott, C.A., Peterson, P.A., Teyton, L., Wilson, I.A.(1998) Immunity 8: 319-329
- PubMed: 9529149
- DOI: https://doi.org/10.1016/s1074-7613(00)80537-3
- Primary Citation of Related Structures:
1IAO, 2IAD - PubMed Abstract:
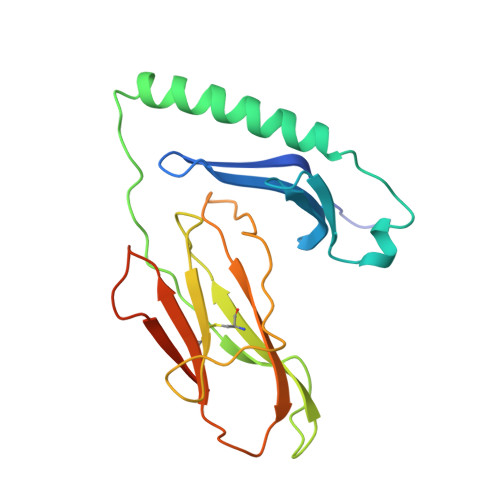
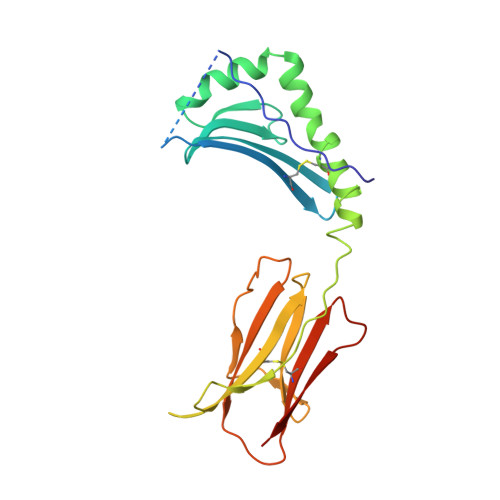
We have determined the structures of I-Ad covalently linked to an ovalbumin peptide (OVA323-339) and to an influenza virus hemagglutinin peptide (HA126-138). The floor of the peptide-binding groove contains an unusual beta bulge, not seen in I-E and DR structures, that affects numerous interactions between the alpha and beta chains and bound peptide. Unlike other MHC-peptide complexes, the peptides do not insert any large anchor residues into the binding pockets of the shallow I-Ad binding groove. The previously identified six-residue "core" binding motif of I-Ad occupies only the P4 to P9 pockets, implying that specificity of T cell receptor recognition of I-Ad-peptide complexes can be accomplished by peptides that only partially fill the MHC groove.
Organizational Affiliation:
Department of Molecular Biology, The Scripps Research Institute, La Jolla, California 92037, USA.