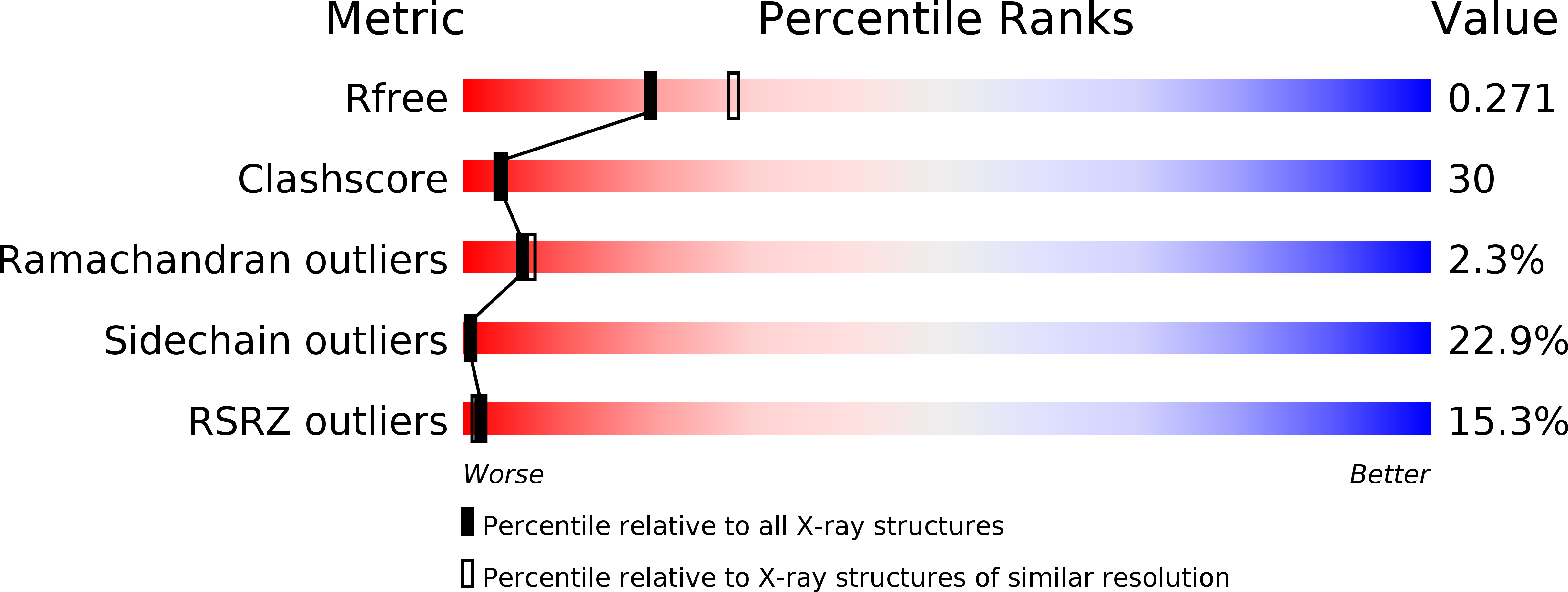
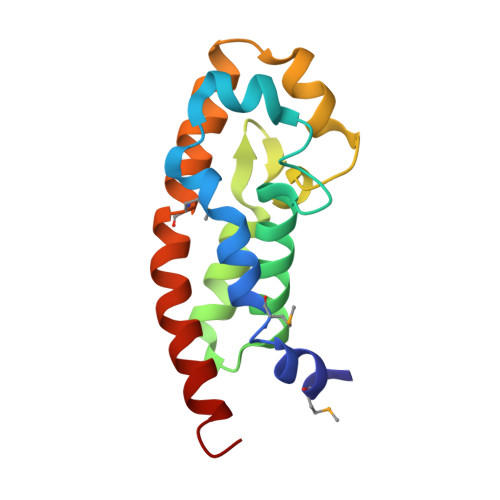
Structure of the Exos Gtpase Activating Domain
Wurtele, M., Renault, L., Barbieri, J.T., Wittinghofer, A., Wolf, E.(2001) FEBS Lett 491: 26
- PubMed: 11226412
- DOI: https://doi.org/10.1016/s0014-5793(01)02105-6
- Primary Citation of Related Structures:
1HE9 - PubMed Abstract:
Pseudomonas aeruginosa is an opportunistic bacterial pathogen of great medical relevance. One of its major toxins, exoenzyme S (ExoS), is a dual function protein with a C-terminal Ras-ADP-ribosylation domain and an N-terminal GTPase activating protein (GAP) domain specific for Rho-family proteins. We report here the three-dimensional structure of the N-terminal domain of ExoS determined by X-ray crystallography to 2.4 A resolution. Its fold is all helical with a four helix bundle core capped by additional irregular helices. Loops that are known to interact with Rho-family proteins show very large mobility. Considering the importance of ExoS in Pseudomonas pathogenicity, this structure could be of interest for drug targeting.
Organizational Affiliation:
Max-Planck-Institut für molekulare Physiologie, Abteilung Strukturelle Biologie, Otto-Hahn-Str. 11, 44227 Dortmund, Germany.