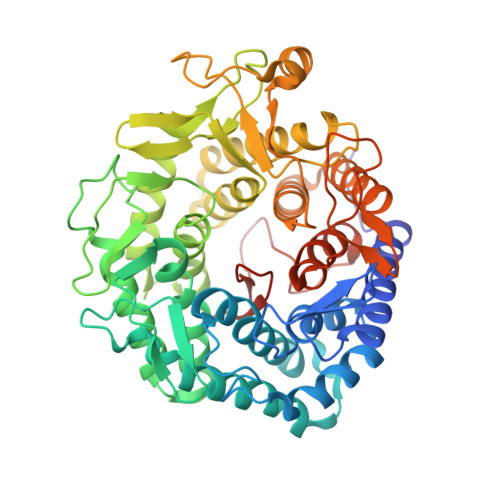
Trichoderma Reesei Alpha-1,2-Mannosidase: Structural Basis for the Cleavage of Four Consecutive Mannose Residues
Van Petegem, F., Contreras, H., Contreras, R., Van Beeumen, J.(2001) J Mol Biol 312: 157
- PubMed: 11545593
- DOI: https://doi.org/10.1006/jmbi.2001.4946
- Primary Citation of Related Structures:
1HCU - PubMed Abstract:
The process of N-glycosylation of eukaryotic proteins involves a range of host enzymes that delete or add saccharide monomers. While endoplasmic reticulum (E.R.) mannosidases cleave only one mannose to produce the Man8B isomer, an alpha-1,2-mannosidase from Trichoderma reesei can sequentially cleave all four 1,2-linked mannose sugars from a Man(9)GlcNAc(2) oligosaccharide, a feature reminiscent of the activity of Golgi mannosidases. We now report the structure of the T. reesei enzyme at 2.37 A resolution. The enzyme folds as an (alpha alpha)(7) barrel. The substrate-binding site of the T. reesei mannosidase differs appreciably from the Saccharomyces cerevisiae enzyme. In the former, shorter loops at the surface allow substrate protein to come closer to the catalytic site. There is more internal space available, so that different oligosaccharide conformations are sterically allowed in the T. reesei alpha-1,2-mannosidase.
Organizational Affiliation:
Department of Biochemistry, Physiology and Microbiology, University of Ghent, K.L. Ledeganckstraat 35, 9000 Ghent, Belgium.