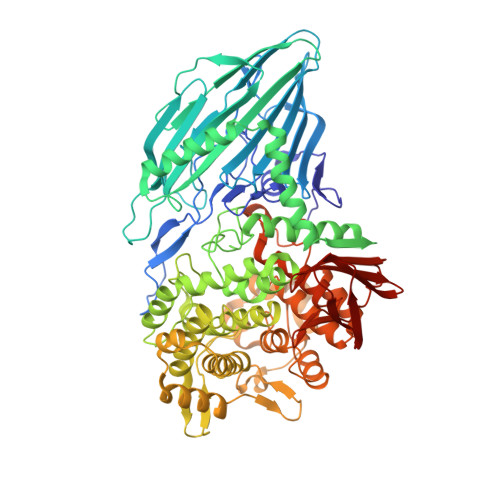
Crystal Structure of Maltose Phosphorylase from Lactobacillus Brevis: Unexpected Evolutionary Relationship with Glucoamylases.
Egloff, M.-P., Uppenberg, J., Haalck, L., Van Tilbeurgh, H.(2001) Structure 9: 689
- PubMed: 11587643
- DOI: https://doi.org/10.1016/s0969-2126(01)00626-8
- Primary Citation of Related Structures:
1H54 - PubMed Abstract:
Maltose phosphorylase (MP) is a dimeric enzyme that catalyzes the conversion of maltose and inorganic phosphate into beta-D-glucose-1-phosphate and glucose without requiring any cofactors, such as pyridoxal phosphate. The enzyme is part of operons that are involved in maltose/malto-oligosaccharide metabolism. Maltose phosphorylases have been classified in family 65 of the glycoside hydrolases. No structure is available for any member of this family. We report here the 2.15 A resolution crystal structure of the MP from Lactobacillus brevis in complex with the cosubstrate phosphate. This represents the first structure of a disaccharide phosphorylase. The structure consists of an N-terminal complex beta sandwich domain, a helical linker, an (alpha/alpha)6 barrel catalytic domain, and a C-terminal beta sheet domain. The (alpha/alpha)6 barrel has an unexpected strong structural and functional analogy with the catalytic domain of glucoamylase from Aspergillus awamori. The only conserved glutamate of MP (Glu487) superposes onto the catalytic residue Glu179 of glucoamylase and likely represents the general acid catalyst. The phosphate ion is bound in a pocket facing the carboxylate of Glu487 and is ideally positioned for nucleophilic attack of the anomeric carbon atom. This site is occupied by the catalytic base carboxylate in glucoamylase. These observations strongly suggest that maltose phosphorylase has evolved from glucoamylase. MP has probably conserved one carboxylate group for acid catalysis and has exchanged the catalytic base for a phosphate binding pocket. The relative positions of the acid catalytic group and the bound phosphate are compatible with a direct-attack mechanism of a glycosidic bond by phosphate, in accordance with inversion of configuration at the anomeric carbon as observed for this enzyme.
Organizational Affiliation:
Architecture et Fonction des Macromolécules Biologiques, Centre National de la Recherche Scientifique, Unite Mixte de Recherche 6098, Université d'Aix-Marseille, I et II, Case 925, 13288, Marseille, France.