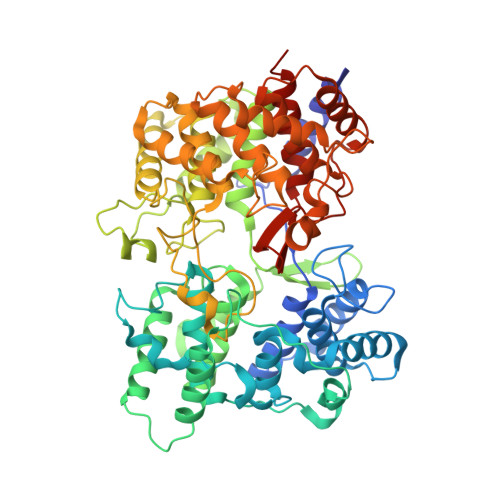
Crystal Structure of a Squalene Cyclase in Complex with the Potential Anticholesteremic Drug Ro48-8071
Lenhart, A., Weihofen, W.A., Pleschke, A.E.W., Schulz, G.E.(2002) Chem Biol 9: 639
- PubMed: 12031670
- DOI: https://doi.org/10.1016/s1074-5521(02)00138-2
- Primary Citation of Related Structures:
1GSZ - PubMed Abstract:
Squalene-hopene cyclase (SHC) catalyzes the conversion of squalene into pentacyclic compounds. It is the prokaryotic counterpart of the eukaryotic oxidosqualene cyclase (OSC) that catalyzes the steroid scaffold formation. Because of clear sequence homology, SHC can serve as a model for OSC, which is an attractive target for anticholesteremic drugs. We have established the crystal structure of SHC complexed with Ro48-8071, a potent inhibitor of OSC and therefore of cholesterol biosynthesis. Ro48-8071 is bound in the active-center cavity of SHC and extends into the channel that connects the cavity with the membrane. The binding site of Ro48-8071 is largely identical with the expected site of squalene; it differs from a previous model based on photoaffinity labeling. The knowledge of the inhibitor binding mode in SHC is likely to help develop more potent inhibitors for OSC.
Organizational Affiliation:
Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, D-79104-, Freiburg im Breisgau, Germany.