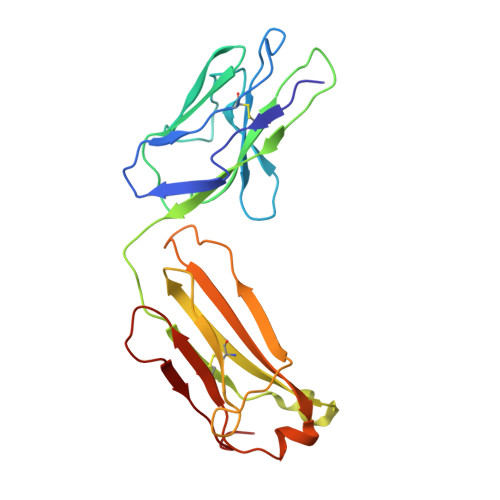
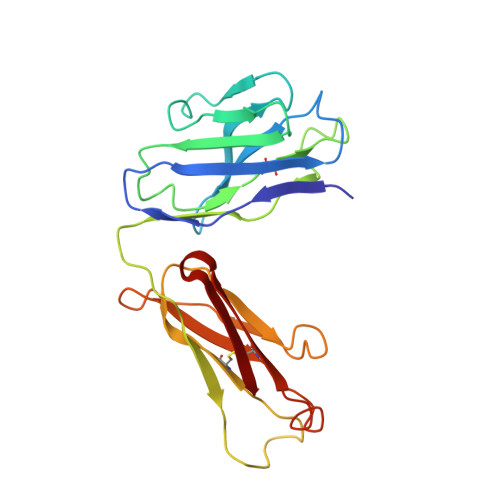
Routes to catalysis: structure of a catalytic antibody and comparison with its natural counterpart.
Haynes, M.R., Stura, E.A., Hilvert, D., Wilson, I.A.(1994) Science 263: 646-652
- PubMed: 8303271
- DOI: https://doi.org/10.1126/science.8303271
- Primary Citation of Related Structures:
1FIG - PubMed Abstract:
The three-dimensional structure of a catalytic antibody (1F7) with chorismate mutase activity has been determined to 3.0 A resolution as a complex with a transition state analog. The structural data suggest that the antibody stabilizes the same conformationally restricted pericyclic transition state as occurs in the uncatalyzed reaction. Overall shape and charge complementarity between the combining site and the transition state analog dictate preferential binding of the correct substrate enantiomer in a conformation appropriate for reaction. Comparison with the structure of a chorismate mutase enzyme indicates an overall similarity between the catalytic mechanism employed by the two proteins. Differences in the number of specific interactions available for restricting the rotational degrees of freedom in the transition state, and the lack of multiple electrostatic interactions that might stabilize charge separation in this highly polarized metastable species, are likely to account for the observed 10(4) times lower activity of the antibody relative to that of the natural enzymes that catalyze this reaction. The structure of the 1F7 Fab'-hapten complex provides confirmation that the properties of an antibody catalyst faithfully reflect the design of the transition state analog.
Organizational Affiliation:
Department of Molecular Biology, Scripps Research Institute, La Jolla, CA 92037.