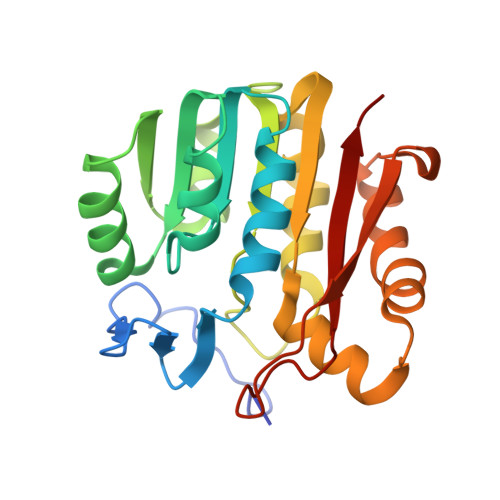
Catalytic mechanism of guanidinoacetate methyltransferase: crystal structures of guanidinoacetate methyltransferase ternary complexes.
Komoto, J., Yamada, T., Takata, Y., Konishi, K., Ogawa, H., Gomi, T., Fujioka, M., Takusagawa, F.(2004) Biochemistry 43: 14385-14394
- PubMed: 15533043
- DOI: https://doi.org/10.1021/bi0486785
- Primary Citation of Related Structures:
1XCJ, 1XCL - PubMed Abstract:
Guanidinoacetate methyltransferase (GAMT) is the enzyme that catalyzes the last step of creatine biosynthesis. The enzyme is found in abundance in the livers of all vertebrates. The intact GAMT from recombinant rat liver has been crystallized with an inhibitor S-adenosylhomocysteine (SAH) and a substrate guanidinoacetate (GAA), and with SAH and an inhibitor guanidine (GUN). These ternary complex structures have been determined at 2.0 A resolution. GAMT has an alpha/beta open-sandwich structure, and the N-terminal section (residues 1-42) covers the active site entrance so that the active site is not visible. SAH has extensive interactions with GAMT through H-bonds and hydrophobic interactions. The guanidino groups of GAA and GUN form two pairs of H-bonds with E45 and D134, respectively. The carboxylate group of GAA interacts with the backbone amide groups of L170 and T171. A model structure of GAMT containing the two substrates (SAM and GAA) was built by attaching a methyl group (C(E)) on S(D) of the bound SAH. On the basis of this model structure, a catalytic mechanism of GAMT is proposed. The active site entrance is opened when the N-terminal section is moved out. GAA and SAM enter the active site and interact with the amino acid residues on the surface of the active site by polar and nonpolar interactions. O(D1) of D134 and C(E) of SAM approach N(E) of GAA from the tetrahedral directions. The O(D1)...N(E) and C(E)...N(E) distances are 2.9 and 2.2 A, respectively. It is proposed that three slightly negatively charged carbonyl oxygen atoms (O of T135, O of C168, and O(B) of GAA) around O(D1) of D134 increase the pK(a) of O(D1) so that O(D1) abstracts the proton on N(E). A strong nucleophile is generated on the deprotonated N(E) of GAA, which abstracts the methyl group (C(E)) from the positively charged S(D) of SAM, and creatine (methyl-GAA) and SAH (demethyl-SAM) are produced. E45, D134, and Y221 mutagenesis studies support the proposed mechanism. A mutagenesis study and the inhibitory mechanism of guanidine analogues support the proposed mechanism.
Organizational Affiliation:
Department of Molecular Biosciences, University of Kansas, 1200 Sunnyside Avenue, Lawrence, Kansas 66045-7534, USA.