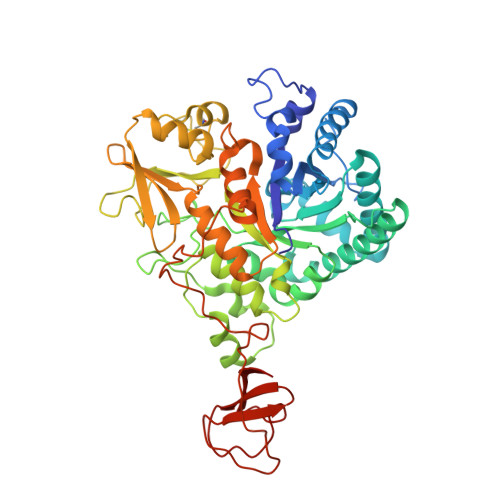
Structure-Based Exploration of Cyclic Dipeptide Chitinase Inhibitors
Houston, D.R., Synstad, B., Eijsink, V.G.H., Stark, M.J., Eggleston, I., Van Aalten, D.M.F.(2004) J Med Chem 47: 5713
- PubMed: 15509170
- DOI: https://doi.org/10.1021/jm049940a
- Primary Citation of Related Structures:
1W1P, 1W1T, 1W1V, 1W1Y - PubMed Abstract:
Family 18 chitinases play an essential role in a range of pathogens and pests. Several inhibitors are known, including the potent inhibitors argadin and allosamidin, and the structures of these in complex with chitinases have been elucidated. Recent structural analysis has revealed that CI-4 [cyclo-(L-Arg-D-Pro)] inhibits family 18 chitinases by mimicking the structure of the proposed reaction intermediate. Here we report the high-resolution structures of four new CI-4 derivatives, cyclo-(L-Arg-L-Pro), cyclo-(Gly-L-Pro), cyclo-(L-His-L-Pro), and cyclo-(L-Tyr-L-Pro), in complex with a family 18 chitinase. In addition, details of enzyme inhibition and in vivo activity against Saccharomyces cerevisiae are presented. The structures reveal that the common cyclo-(Gly-Pro) substructure is sufficient for binding, allowing modification of the side chain of the nonproline residue. This suggests that design of cyclic dipeptides with a view to increasing inhibition of family 18 chitinases should be possible through relatively accessible chemistry. The derivatives presented here in complex with chitinase B from Serratia marcescens provide further insight into the mechanism of inhibition of chitinases by cyclic dipeptides as well as providing a new scaffold for chitinase inhibitor design.
Organizational Affiliation:
Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland, UK.