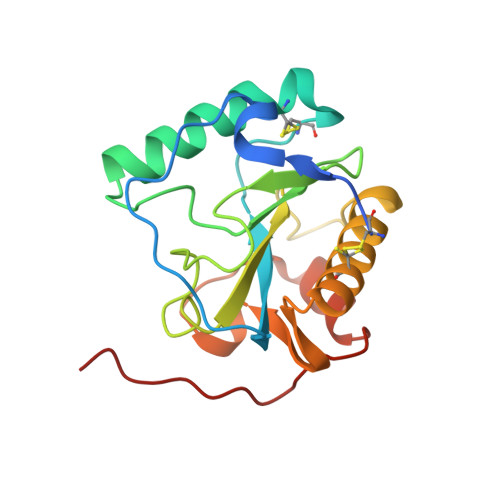
Crystal structure of the Drosophila peptidoglycan recognition protein (PGRP)-SA at 1.56 A resolution
Reiser, J.B., Teyton, L., Wilson, I.A.(2004) J Mol Biol 340: 909-917
- PubMed: 15223330
- DOI: https://doi.org/10.1016/j.jmb.2004.04.077
- Primary Citation of Related Structures:
1SXR - PubMed Abstract:
Peptidoglycan recognition proteins (PGRPs) form a recently discovered protein family, which is conserved from insect to mammals and is implicated in the innate immune system by interacting with/or degrading microbial peptidoglycans (PGNs). Drosophila PGRP-SA is a member of this family of pattern recognition receptors and is involved in insect Toll activation. We report here the crystal structure of PGRP-SA at 1.56 A resolution, which represents the first example of a "recognition" PGRP. Comparison with the catalytic Drosophila PGRP-LB reveals an overall structure conservation with an L-shaped hydrophilic groove that is likely the PGN carbohydrate core binding site, but further suggests some possible functional homology between recognition and catalytic PGRPs. Consistent with sequence analysis, PGRP-SA does not contain the canonical zinc-binding residues found in catalytic PGRPs. However, substitution of the zinc-binding cysteine residue by serine, along with an altered coordinating histidine residue, assembles a constellation of residues that resembles a modified catalytic triad. The serine/histidine juxtaposition to a threonine residue and a carbonyl oxygen atom, along with conservation of the catalytic water molecule found in PGRP-LB, tantalizingly suggests some hydrolytic function for this member of receptor PGRPs.
Organizational Affiliation:
Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla CA 92037, USA.