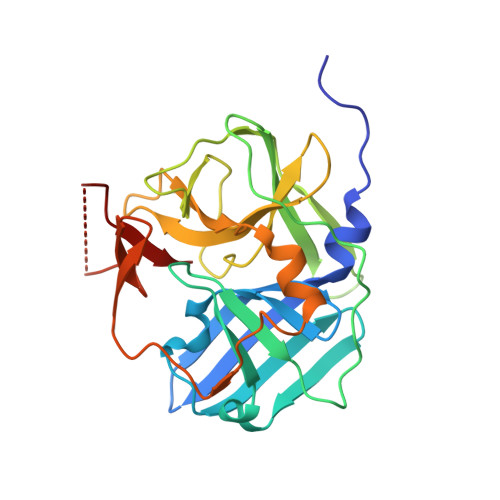
Crystal structure of tobacco etch virus protease shows the protein C terminus bound within the active site.
Nunn, C.M., Jeeves, M., Cliff, M.J., Urquhart, G.T., George, R.R., Chao, L.H., Tscuchia, Y., Djordjevic, S.(2005) J Mol Biol 350: 145-155
- PubMed: 15919091
- DOI: https://doi.org/10.1016/j.jmb.2005.04.013
- Primary Citation of Related Structures:
1Q31 - PubMed Abstract:
Tobacco etch virus (TEV) protease is a cysteine protease exhibiting stringent sequence specificity. The enzyme is widely used in biotechnology for the removal of the affinity tags from recombinant fusion proteins. Crystal structures of two TEV protease mutants as complexes with a substrate and a product peptide provided the first insight into the mechanism of substrate specificity of this enzyme. We now report a 2.7A crystal structure of a full-length inactive C151A mutant protein crystallised in the absence of peptide. The structure reveals the C terminus of the protease bound to the active site. In addition, we determined dissociation constants of TEV protease substrate and product peptides using isothermal titration calorimetry for various forms of this enzyme. Data suggest that TEV protease could be inhibited by the peptide product of autolysis. Separate modes of recognition for native substrates and the site of TEV protease self-cleavage are proposed.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, University College London, Gower Street, London, WC1E 6BT, UK.