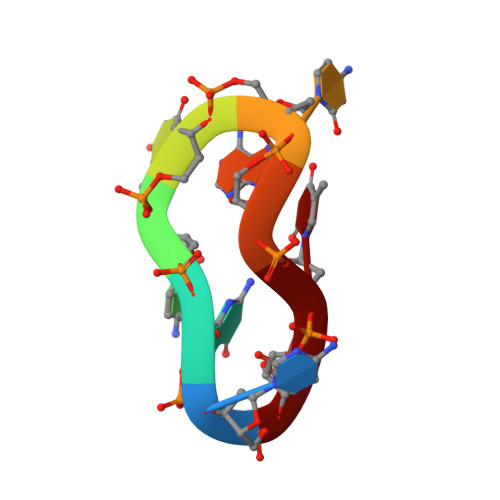Four-stranded DNA structure stabilized by a novel G:C:A:T tetrad.
Escaja, N., Gelpi, J.L., Orozco, M., Rico, M., Pedroso, E., Gonzalez, C.(2003) J Am Chem Soc 125: 5654-5662
- PubMed: 12733903
- DOI: https://doi.org/10.1021/ja0344157
- Primary Citation of Related Structures:
1N96 - PubMed Abstract:
The solution structure of a cyclic oligonucleotide d
has been determined by two-dimensional NMR spectroscopy and restrained molecular dynamics. Under the appropriate experimental conditions, this molecule self-associates, forming a symmetric dimer stabilized by four intermolecular Watson-Crick base pairs. The resulting four-stranded structure consists of two G:C:A:T tetrads, formed by facing the minor groove side of the Watson-Crick base-pairs. Most probably, the association of the base-pairs is stabilized by coordinating a Na(+) cation. This is the first time that this novel G:C:A:T tetrad has been found in an oligonucleotide structure. This observation increases considerably the number of sequences that may adopt a four-stranded architecture. Overall, the three-dimensional structure is similar to those observed previously in other quadruplexes formed by minor groove alignment of Watson-Crick base pairs. This resemblance strongly suggests that we may be observing a general motif for DNA-DNA recognition.
Organizational Affiliation:
Departament de Química Orgànica, Universitat de Barcelona, C/, Martí i Franquès 1-11, 08028 Barcelona, Spain.