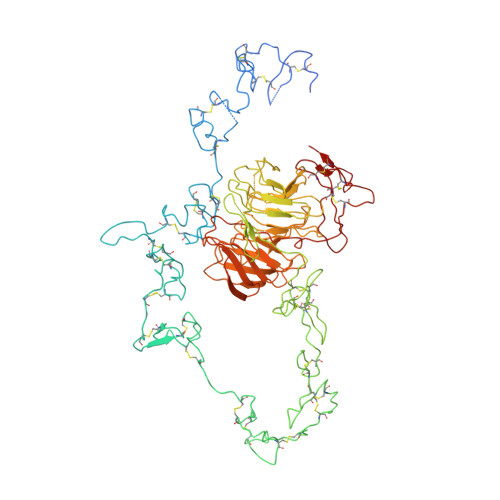
Structure of the LDL receptor extracellular domain at endosomal pH
Rudenko, G., Henry, L., Henderson, K., Ichtchenko, K., Brown, M.S., Goldstein, J.L., Deisenhofer, J.(2002) Science 298: 2353-2358
- PubMed: 12459547
- DOI: https://doi.org/10.1126/science.1078124
- Primary Citation of Related Structures:
1N7D - PubMed Abstract:
The low-density lipoprotein receptor mediates cholesterol homeostasis through endocytosis of lipoproteins. It discharges its ligand in the endosome at pH < 6. In the crystal structure at pH = 5.3, the ligand-binding domain (modules R2 to R7) folds back as an arc over the epidermal growth factor precursor homology domain (the modules A, B, beta propeller, and C). The modules R4 and R5, which are critical for lipoprotein binding, associate with the beta propeller via their calcium-binding loop. We propose a mechanism for lipoprotein release in the endosome whereby the beta propeller functions as an alternate substrate for the ligand-binding domain, binding in a calcium-dependent way and promoting lipoprotein release.
Organizational Affiliation:
Department of Biochemistry, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard Y4-206, Dallas, TX 75390, USA.