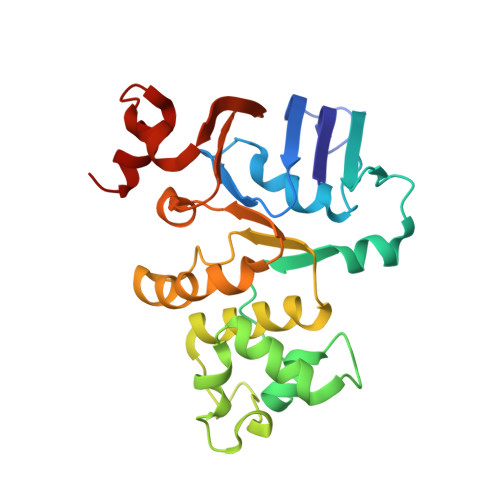
Crystal structure of the nucleotide binding domain of the ABC-transporter hemolysin B: identification of a variable region within ABC helical domains
Schmitt, L., Benabdelhak, H., Blight, M.A., Holland, I.B., Stubbs, M.T.(2003) J Mol Biol 330: 333-342
- PubMed: 12823972
- DOI: https://doi.org/10.1016/s0022-2836(03)00592-8
- Primary Citation of Related Structures:
1MT0 - PubMed Abstract:
The ABC-transporter haemolysin B is a central component of the secretion machinery that translocates the toxin, haemolysin A, in a Sec-independent fashion across both membranes of E. coli. Here, we report the X-ray crystal structure of the nucleotide-binding domain (NBD) of HlyB. The molecule shares the common overall architecture of ABC-transporter NBDs. However, the last three residues of the Walker A motif adopt a 3(10) helical conformation, stabilized by a bound anion. In consequence, this results in an unusual interaction between the Walker A lysine residue and the Walker B glutamate residue. As these residues are normally required to be available for ATP binding, for catalysis and for dimer formation of ABC domains, we suggest that this conformation may represent a latent monomeric form of the NBD. Surprisingly, comparison of available NBD structures revealed a structurally diverse region (SDR) of about 30 residues within the helical arm II domain, unique to each of the eight NBDs analyzed. As this region interacts with the transmembrane part of ABC-transporters, the SDR helps to explain the selectivity and/or targeting of different NBDs to their cognate transmembrane domains.
Organizational Affiliation:
Institut für Biochemie, Biozentrum N210, Johann Wolfgang Goethe Universität Frankfurt, Marie-Curie Str. 9, 60439, Frankfurt, Germany. lschmitt@em.uni-frankfurt.de