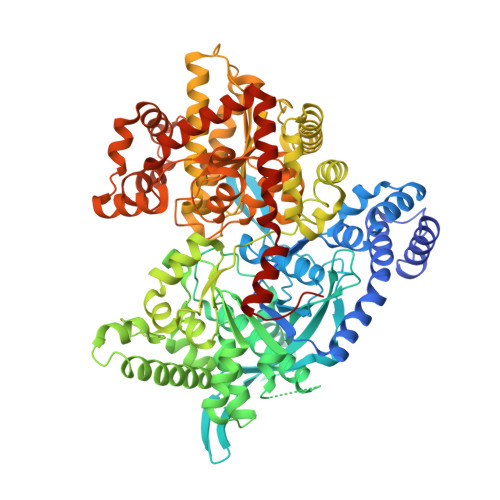
Human liver glycogen phosphorylase inhibitors bind at a new allosteric site.
Rath, V.L., Ammirati, M., Danley, D.E., Ekstrom, J.L., Gibbs, E.M., Hynes, T.R., Mathiowetz, A.M., McPherson, R.K., Olson, T.V., Treadway, J.L., Hoover, D.J.(2000) Chem Biol 7: 677-682
- PubMed: 10980448
- DOI: https://doi.org/10.1016/s1074-5521(00)00004-1
- Primary Citation of Related Structures:
1EM6, 1EXV - PubMed Abstract:
Glycogen phosphorylases catalyze the breakdown of glycogen to glucose-1-phosphate for glycolysis. Maintaining control of blood glucose levels is critical in minimizing the debilitating effects of diabetes, making liver glycogen phosphorylase a potential therapeutic target. The binding site in human liver glycogen phosphorylase (HLGP) for a class of promising antidiabetic agents was identified crystallographically. The site is novel and functions allosterically by stabilizing the inactive conformation of HLGP. The initial view of the complex revealed key structural information and inspired the design of a new class of inhibitors which bind with nanomolar affinity and whose crystal structure is also described. We have identified the binding site of a new class of allosteric HLGP inhibitors. The crystal structure revealed the details of inhibitor binding, led to the design of a new class of compounds, and should accelerate efforts to develop therapeutically relevant molecules for the treatment of diabetes.
Organizational Affiliation:
Department of Exploratory Medicinal Sciences, Global Research and Development, Pfizer Inc., Groton, CT 06340, USA.