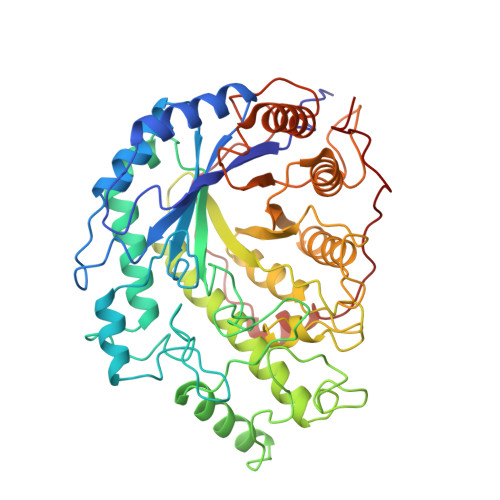
Crystal structures of soybean beta-amylase reacted with beta-maltose and maltal: active site components and their apparent roles in catalysis.
Mikami, B., Degano, M., Hehre, E.J., Sacchettini, J.C.(1994) Biochemistry 33: 7779-7787
- PubMed: 8011643
- Primary Citation of Related Structures:
1BYA, 1BYB, 1BYC, 1BYD - PubMed Abstract:
The crystal structures of catalytically competent soybean beta-amylase, unliganded and bathed with small substrates (beta-maltose, maltal), were determined at 1.9-2.2-A resolution. Two molecules of beta-maltose substrate bind to the protein in tandem, with some maltotetraose enzymic condensation product sharing the same binding sites. The beta-amylase soaked with maltal shows a similar arrangement of two bound molecules of 2-deoxymaltose, the enzymic hydration product. In each case the nonreducing ends of the saccharide ligands are oriented toward the base of the protein's active site pocket. The catalytic center, located between the bound disaccharides and found deeper in the pocket than where the inhibitor alpha-cyclodextrin binds, is characterized by the presence of oppositely disposed carboxyl groups of two conserved glutamic acid residues. The OE2 carboxyl of Glu 186 is below the plane of the penultimate glucose residue (Glc 2) of bound maltotetraose, 2.6 A from the oxygen atom of that ligand's penultimate alpha-1,4-glucosidic linkage. The OE2 carboxyl of Glu 380 lies above the plane of Glc 2, 2.8 A from the O-1 atom of the more deeply bound beta-maltose. Saccharide binding does not alter the spatial coordinates of these two carboxyl groups or the overall conformation of the 57-kDa protein. However, the saccharide complexes of the active enzyme are associated with a significant (10 A) local conformational change in a peptide segment of a loop (L3) that borders the active site pocket.(ABSTRACT TRUNCATED AT 250 WORDS)
Organizational Affiliation:
Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York 10461.