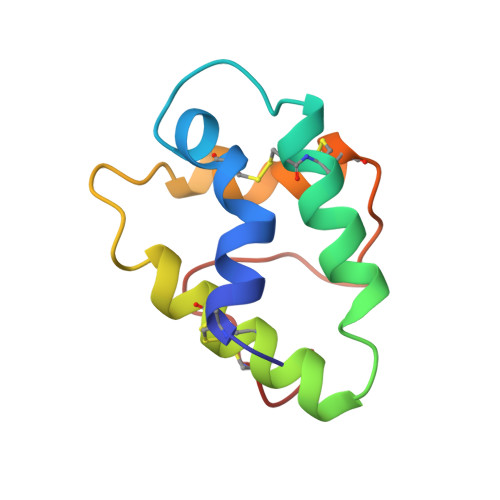
The crystal structure of a wheat nonspecific lipid transfer protein (ns-LTP1) complexed with two molecules of phospholipid at 2.1 A resolution.
Charvolin, D., Douliez, J.P., Marion, D., Cohen-Addad, C., Pebay-Peyroula, E.(1999) Eur J Biochem 264: 562-568
- PubMed: 10491104
- DOI: https://doi.org/10.1046/j.1432-1327.1999.00667.x
- Primary Citation of Related Structures:
1BWO - PubMed Abstract:
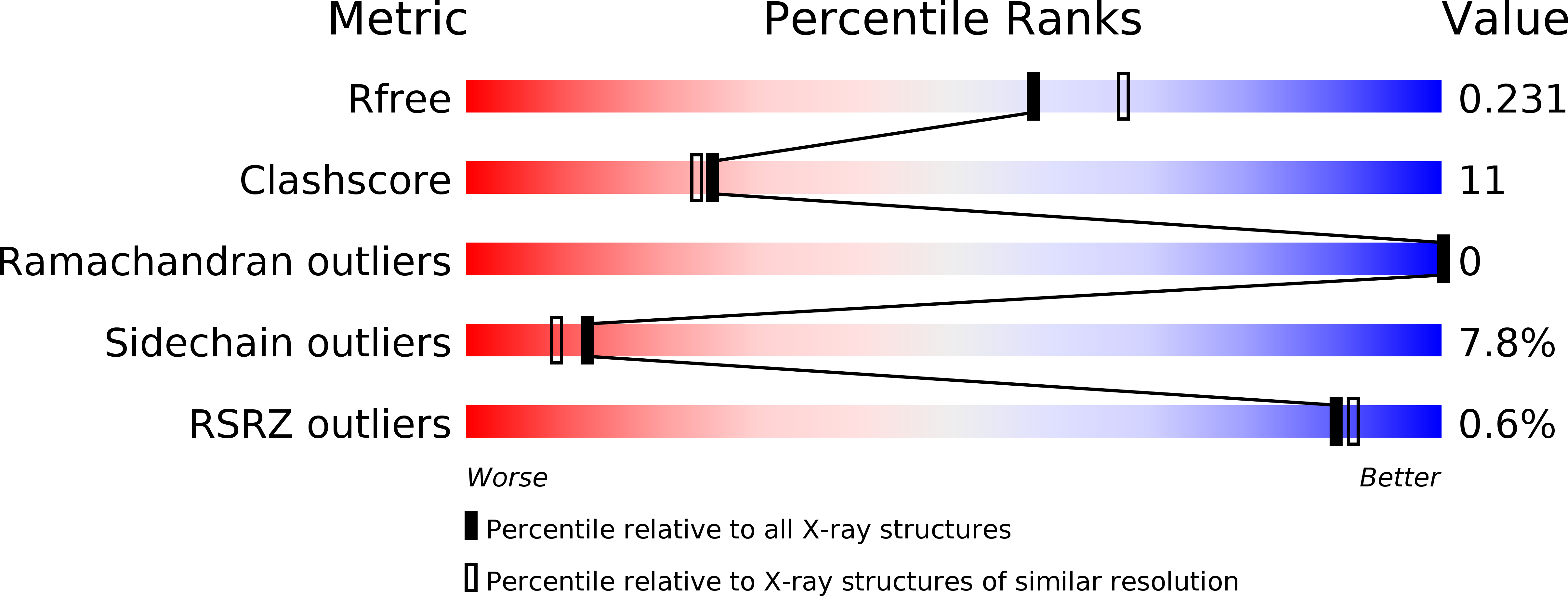
Nonspecific lipid transfer proteins (ns-LTP1) form a multigenic protein family in plants. In vitro they are able to bind all sort of lipids but their function, in vivo, remains speculative. A ns-LTP1 isolated from wheat seed was crystallized in the presence of lyso-myristoyl-phosphatidylcholine (LMPC). The structure was solved by molecular replacement and refined to 2.1 A resolution to an R-factor of 16.3% and a free R-factor of 21.3%. It reveals for the first time that the protein binds two LMPC molecules that are inserted head to tail in a hydrophobic cavity. A detailed study of the structure leads to the conclusion that there are two lipid-binding sites, one of which shows a higher affinity for the LMPC than the other. Comparison with other structures of lipid-bound ns-LTP1 suggests that the presence of two binding sites is a general feature of plant ns-LTP1.
Organizational Affiliation:
Institut de Biologie Structural Jean-Pierre Ebel, CEA-CNRS, Nantes, France.