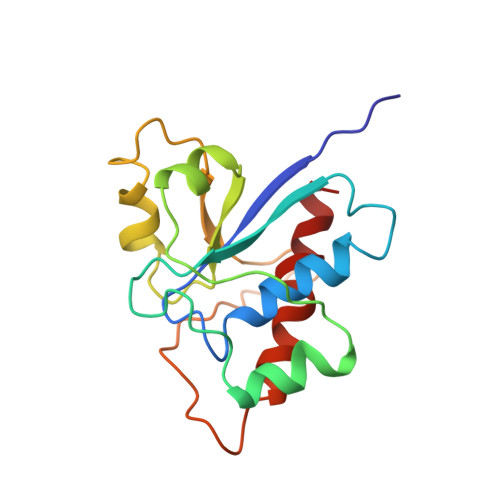
Solution structure of a low molecular weight protein tyrosine phosphatase.
Logan, T.M., Zhou, M.M., Nettesheim, D.G., Meadows, R.P., Van Etten, R.L., Fesik, S.W.(1994) Biochemistry 33: 11087-11096
- PubMed: 7727361
- DOI: https://doi.org/10.1021/bi00203a005
- Primary Citation of Related Structures:
1BVH - PubMed Abstract:
Protein tyrosine phosphatases (PTPs) are important enzymes involved in signal transduction, cell cycle regulation, and the control of differentiation. Despite the importance of this class of enzymes in the control of critical cell processes, very little structural information is available for this family of proteins. In this paper, we present the first solution structure of a protein tyrosine phosphatase. This protein is a low molecular weight cytosolic PTP that was initially isolated from bovine heart. The structure that was determined from 1747 NMR-derived restraints consists of a central four-stranded parallel beta-sheet surrounded by four alpha-helices and a short 3(10) helix. The phosphate binding site, identified by chemical shift changes upon the addition of the competitive inhibitors phosphate and vanadate, is in a loop region connecting the C-terminal end of the first beta-strand with the first alpha-helix. Residues in the second, fourth, and fifth alpha-helices and in some of the loop regions connecting the elements of regular secondary structure also contribute to the binding site. The structure determined here is consistent with previous mutagenesis and chemical modification studies conducted on this protein.
Organizational Affiliation:
Pharmaceutical Discovery Division, Abbott Laboratories, Abbott Park, Illinois 60064, USA.