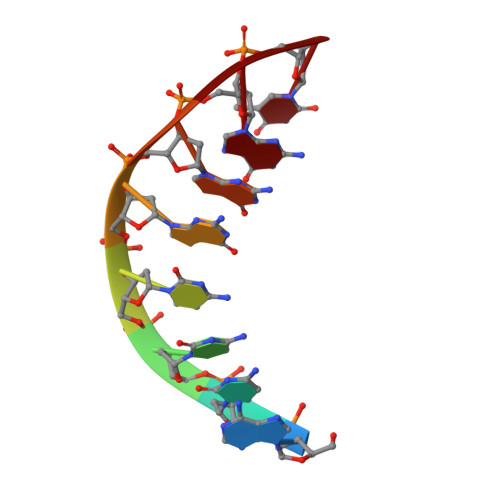
The three-dimensional structure of the 4:1 mithramycin:d(ACCCGGGT)2 complex: evidence for an interaction between the E saccharides
Keniry, M.A., Owen, E.A., Shafer, R.H.(2000) Biopolymers 54: 104-114
- PubMed: 10861371
- DOI: https://doi.org/10.1002/1097-0282(200008)54:2<104::AID-BIP3>3.0.CO;2-2
- Primary Citation of Related Structures:
1BP8 - PubMed Abstract:
Mithramycin and chromomycin, two antitumor drugs, each having an identical aglycone and nearly identical disaccharide and trisaccharide side chains, have differing binding properties to a small oligonucleotide, d(ACCCGGGT)(2) (M. A. Keniry et al., Journal of Molecular Biology, 1993, Vol. 231, pp. 753-767). In order to understand the forces that induce four mithramycin molecules to bind to d(ACCCGGGT)(2) instead of two drug molecules in the case of chromomycin, the structure of the 4:2:1 mithramycin: Mg(2+):d(ACCCGGGT)(2) complex was investigated by (1)H-nmr and restrained molecular dynamics. The resulting three-dimensional model showed that in order to accommodate the close approach of one neighboring mithramycin dimer, the inwardly directed CDE saccharide chain of the neighboring mithramycin dimer undergoes a conformational change such that the E saccharide no longer spans the minor groove but reorients so that the hydrophilic face of the E saccharides from the two dimers oppose each other. Two hydrogen bonds are formed between the hydroxyl groups of the two opposing E saccharide groups. The results are interpreted in terms of the differences in stereochemistry and functional group substitutions between mithramycin and chromomycin. A mithramycin dimer is able to self-associate on an oligonucleotide template because it has two hydroxyl groups on the same face of its terminal E saccharide. A chromomycin dimer is unable to self-associate because one of these hydroxyl groups is acetylated and the neighboring hydroxyl group has a stereochemistry that cannot permit close contact of the hydroxyl group with a neighbouring chromomycin dimer.
Organizational Affiliation:
Research School of Chemistry, The Australian National University, Canberra, ACT 0200, Australia.