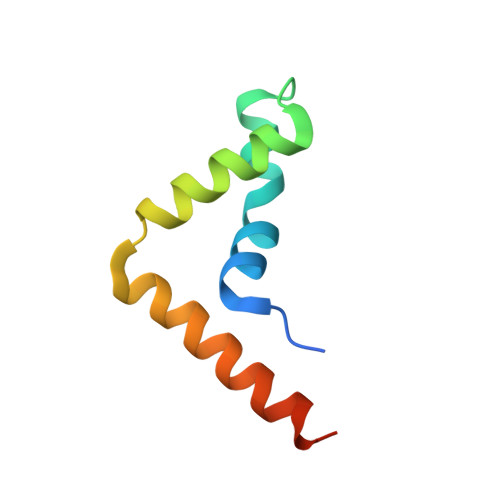
Crystal structure of the HMG domain of human BAF57 and its interaction with four-way junction DNA.
Heo, Y., Park, J.H., Kim, J., Han, J., Yun, J.H., Lee, W.(2020) Biochem Biophys Res Commun 533: 919-924
- PubMed: 33010889
- DOI: https://doi.org/10.1016/j.bbrc.2020.09.094
- Primary Citation of Related Structures:
7CYU - PubMed Abstract:
The SWI/SNF chromatin remodeling complex plays important roles in gene regulation and it is classified as the SWI/SNF complex in yeast and BAF complex in vertebrates. BAF57, one of the subunits that forms the chromatin remodeling complex core, is well conserved in the BAF complex of vertebrates, which is replaced by bap111 in the Drosophila BAP complex and does not have a counterpart in the yeast SWI/SNF complex. This suggests that BAF57 is a key component of the chromatin remodeling complex in higher eukaryotes. BAF57 contains a HMG domain, which is widely distributed among various proteins and functions as a DNA binding motif. Most proteins with HMG domain bind to four-way junction (4WJ) DNA. Here, we report the crystal structure of the HMG domain of BAF57 (BAF57 HMG ) at a resolution of 2.55 Å. The structure consists of three α-helices and adopts an L-shaped form. The overall structure is stabilized by a hydrophobic core, which is formed by hydrophobic residues. The binding affinity between BAF57 HMG and 4WJ DNA is determined as a 295.83 ± 1.05 nM using a fluorescence quenching assay, and the structure revealed 4WJ DNA binding site of BAF57 HMG . Our data will serve structural basis in understanding the roles of BAF57 during chromatin remodeling process.
Organizational Affiliation:
Structural Biochemistry & Molecular Biophysics Laboratory, Department of Biochemistry, College of Life Sciences & Biotechnology, Yonsei University, Seoul, 120-749, South Korea.