Modulation of Amyloidogenic Protein Self-Assembly Using Tethered Small Molecules.
Cawood, E.E., Guthertz, N., Ebo, J.S., Karamanos, T.K., Radford, S.E., Wilson, A.J.(2020) J Am Chem Soc 142: 20845-20854
- PubMed: 33253560
- DOI: https://doi.org/10.1021/jacs.0c10629
- Primary Citation of Related Structures:
7AFV - PubMed Abstract:
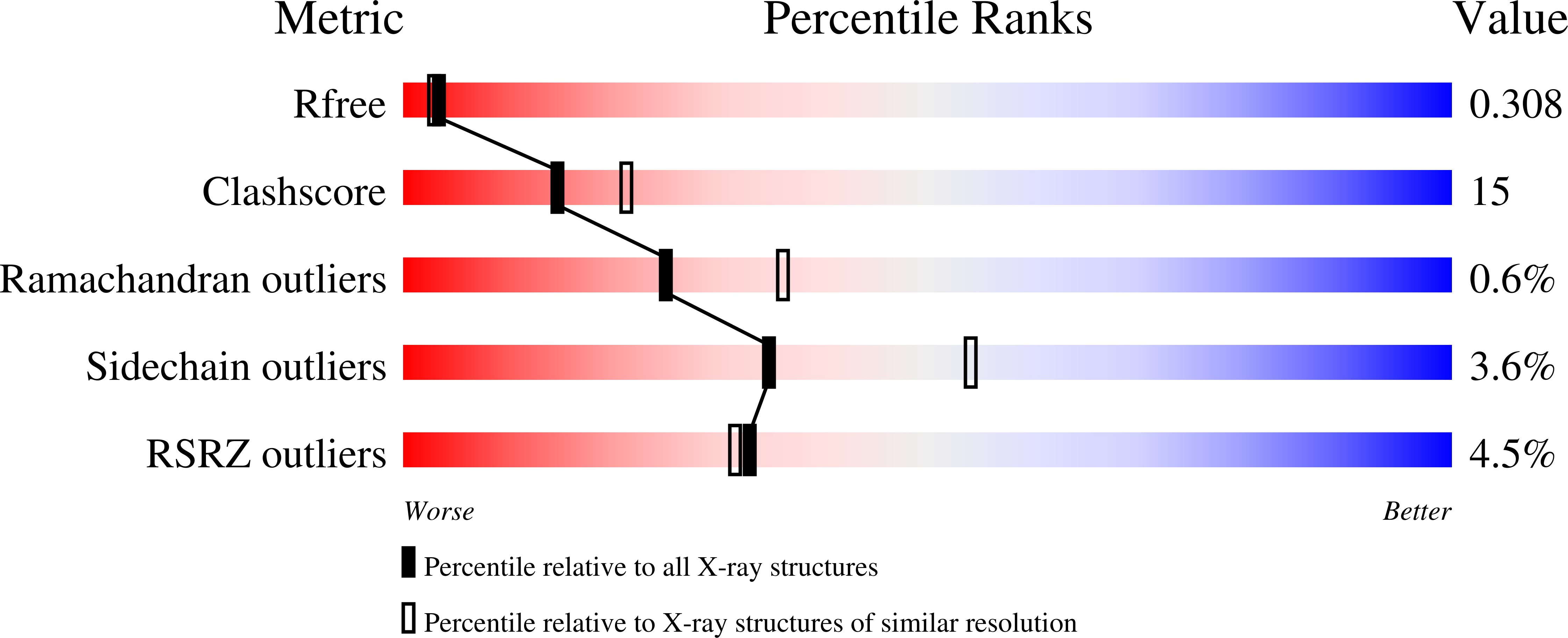
Protein-protein interactions (PPIs) are involved in many of life's essential biological functions yet are also an underlying cause of several human diseases, including amyloidosis. The modulation of PPIs presents opportunities to gain mechanistic insights into amyloid assembly, particularly through the use of methods which can trap specific intermediates for detailed study. Such information can also provide a starting point for drug discovery. Here, we demonstrate that covalently tethered small molecule fragments can be used to stabilize specific oligomers during amyloid fibril formation, facilitating the structural characterization of these assembly intermediates. We exemplify the power of covalent tethering using the naturally occurring truncated variant (ΔN6) of the human protein β 2 -microglobulin (β 2 m), which assembles into amyloid fibrils associated with dialysis-related amyloidosis. Using this approach, we have trapped tetramers formed by ΔN6 under conditions which would normally lead to fibril formation and found that the degree of tetramer stabilization depends on the site of the covalent tether and the nature of the protein-fragment interaction. The covalent protein-ligand linkage enabled structural characterization of these trapped, off-pathway oligomers using X-ray crystallography and NMR, providing insight into why tetramer stabilization inhibits amyloid assembly. Our findings highlight the power of "post-translational chemical modification" as a tool to study biological molecular mechanisms.
Organizational Affiliation:
Astbury Centre for Structural Molecular Biology, University of Leeds, Leeds LS2 9JT, United Kingdom.