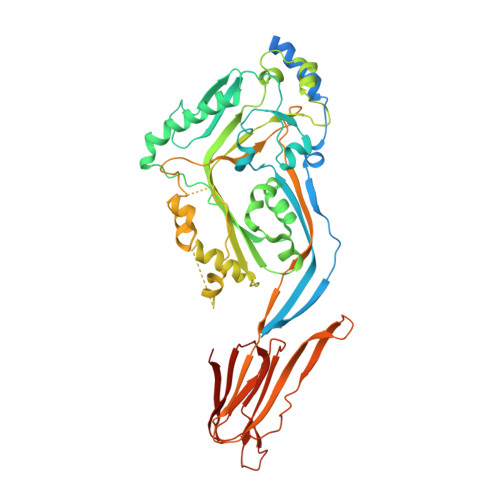
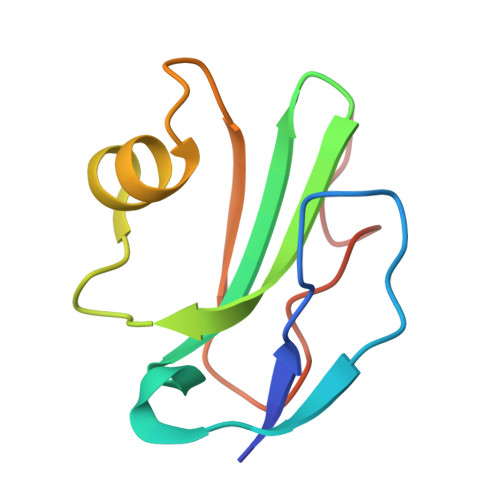
Structural basis for tuning activity and membrane specificity of bacterial cytolysins.
Shah, N.R., Voisin, T.B., Parsons, E.S., Boyd, C.M., Hoogenboom, B.W., Bubeck, D.(2020) Nat Commun 11: 5818-5818
- PubMed: 33199689
- DOI: https://doi.org/10.1038/s41467-020-19482-6
- Primary Citation of Related Structures:
6ZD0 - PubMed Abstract:
Cholesterol-dependent cytolysins (CDCs) are pore-forming proteins that serve as major virulence factors for pathogenic bacteria. They target eukaryotic cells using different mechanisms, but all require the presence of cholesterol to pierce lipid bilayers. How CDCs use cholesterol to selectively lyse cells is essential for understanding virulence strategies of several pathogenic bacteria, and for repurposing CDCs to kill new cellular targets. Here we address that question by trapping an early state of pore formation for the CDC intermedilysin, bound to the human immune receptor CD59 in a nanodisc model membrane. Our cryo electron microscopy map reveals structural transitions required for oligomerization, which include the lateral movement of a key amphipathic helix. We demonstrate that the charge of this helix is crucial for tuning lytic activity of CDCs. Furthermore, we discover modifications that overcome the requirement of cholesterol for membrane rupture, which may facilitate engineering the target-cell specificity of pore-forming proteins.
Organizational Affiliation:
Department of Life Sciences, Sir Ernst Chain Building, Imperial College London, London, SW7 2AZ, UK.