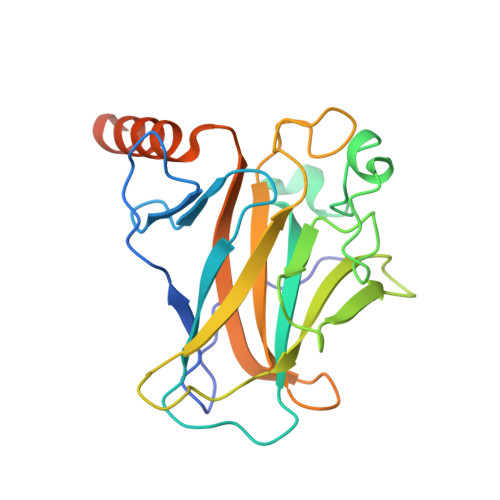
Targeting Cavity-Creating p53 Cancer Mutations with Small-Molecule Stabilizers: the Y220X Paradigm.
Bauer, M.R., Kramer, A., Settanni, G., Jones, R.N., Ni, X., Khan Tareque, R., Fersht, A.R., Spencer, J., Joerger, A.C.(2020) ACS Chem Biol 15: 657-668
- PubMed: 31990523
- DOI: https://doi.org/10.1021/acschembio.9b00748
- Primary Citation of Related Structures:
6SHZ, 6SI0, 6SI1, 6SI2, 6SI3, 6SI4 - PubMed Abstract:
We have previously shown that the thermolabile, cavity-creating p53 cancer mutant Y220C can be reactivated by small-molecule stabilizers. In our ongoing efforts to unearth druggable variants of the p53 mutome, we have now analyzed the effects of other cancer-associated mutations at codon 220 on the structure, stability, and dynamics of the p53 DNA-binding domain (DBD). We found that the oncogenic Y220H, Y220N, and Y220S mutations are also highly destabilizing, suggesting that they are largely unfolded under physiological conditions. A high-resolution crystal structure of the Y220S mutant DBD revealed a mutation-induced surface crevice similar to that of Y220C, whereas the corresponding pocket's accessibility to small molecules was blocked in the structure of the Y220H mutant. Accordingly, a series of carbazole-based small molecules, designed for stabilizing the Y220C mutant, also bound to and stabilized the folded state of the Y220S mutant, albeit with varying affinities due to structural differences in the binding pocket of the two mutants. Some of the compounds also bound to and stabilized the Y220N mutant, but not the Y220H mutant. Our data validate the Y220S and Y220N mutants as druggable targets and provide a framework for the design of Y220S or Y220N-specific compounds as well as compounds with dual Y220C/Y220S specificity for use in personalized cancer therapy.
Organizational Affiliation:
MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge Biomedical Campus, Cambridge CB2 0QH, United Kingdom.