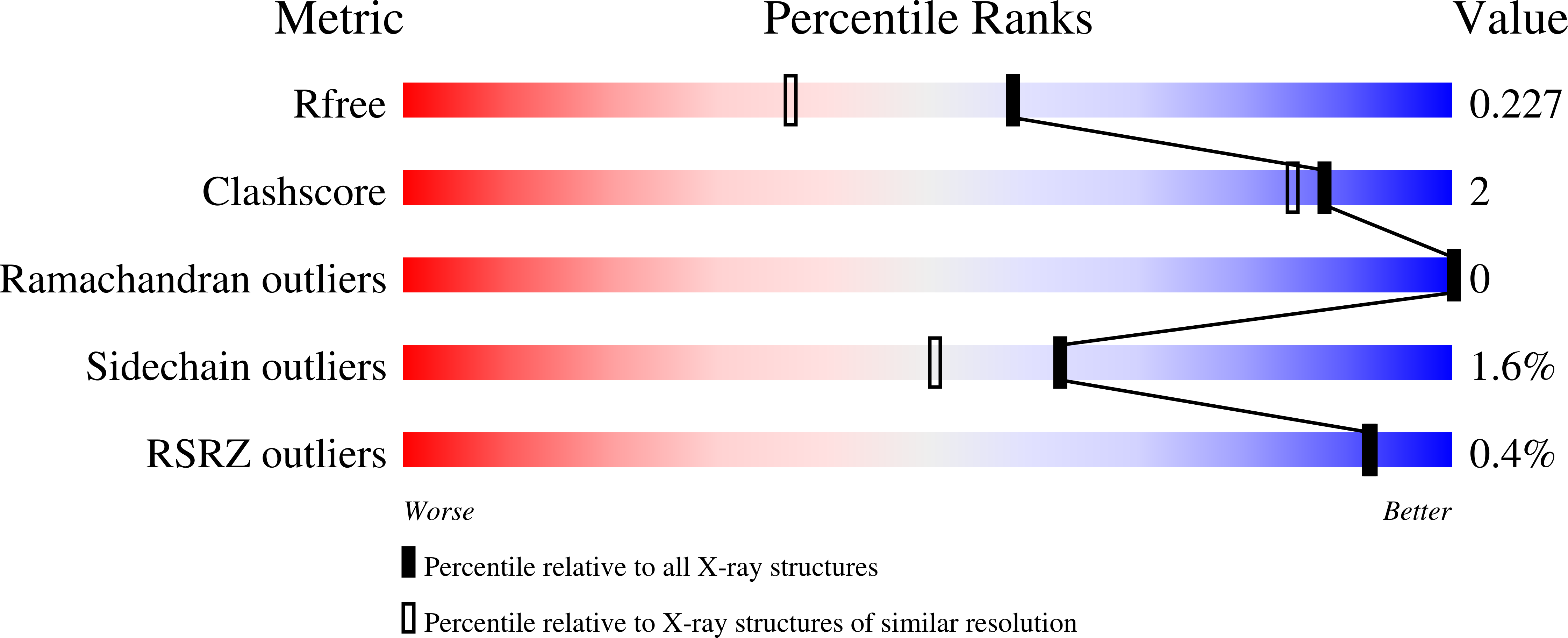
Small-molecule factor B inhibitor for the treatment of complement-mediated diseases.
Schubart, A., Anderson, K., Mainolfi, N., Sellner, H., Ehara, T., Adams, C.M., Mac Sweeney, A., Liao, S.M., Crowley, M., Littlewood-Evans, A., Sarret, S., Wieczorek, G., Perrot, L., Dubost, V., Flandre, T., Zhang, Y., Smith, R.J.H., Risitano, A.M., Karki, R.G., Zhang, C., Valeur, E., Sirockin, F., Gerhartz, B., Erbel, P., Hughes, N., Smith, T.M., Cumin, F., Argikar, U.A., Haraldsson, B., Mogi, M., Sedrani, R., Wiesmann, C., Jaffee, B., Maibaum, J., Flohr, S., Harrison, R., Eder, J.(2019) Proc Natl Acad Sci U S A 116: 7926-7931
- PubMed: 30926668
- DOI: https://doi.org/10.1073/pnas.1820892116
- Primary Citation of Related Structures:
6QSW, 6QSX, 6RAV - PubMed Abstract:
Dysregulation of the alternative complement pathway (AP) predisposes individuals to a number of diseases including paroxysmal nocturnal hemoglobinuria, atypical hemolytic uremic syndrome, and C3 glomerulopathy. Moreover, glomerular Ig deposits can lead to complement-driven nephropathies. Here we describe the discovery of a highly potent, reversible, and selective small-molecule inhibitor of factor B, a serine protease that drives the central amplification loop of the AP. Oral administration of the inhibitor prevents KRN-induced arthritis in mice and is effective upon prophylactic and therapeutic dosing in an experimental model of membranous nephropathy in rats. In addition, inhibition of factor B prevents complement activation in sera from C3 glomerulopathy patients and the hemolysis of human PNH erythrocytes. These data demonstrate the potential therapeutic value of using a factor B inhibitor for systemic treatment of complement-mediated diseases and provide a basis for its clinical development.
Organizational Affiliation:
Novartis Institutes for BioMedical Research, Novartis Pharma AG, CH-4056 Basel, Switzerland.