New Irreversible alpha-l-Iduronidase Inhibitors and Activity-Based Probes.
Artola, M., Kuo, C.L., McMahon, S.A., Oehler, V., Hansen, T., van der Lienden, M., He, X., van den Elst, H., Florea, B.I., Kermode, A.R., van der Marel, G.A., Gloster, T.M., Codee, J.D.C., Overkleeft, H.S., Aerts, J.M.F.G.(2018) Chemistry 24: 19081-19088
- PubMed: 30307091
- DOI: https://doi.org/10.1002/chem.201804662
- Primary Citation of Related Structures:
6I6R, 6I6X - PubMed Abstract:
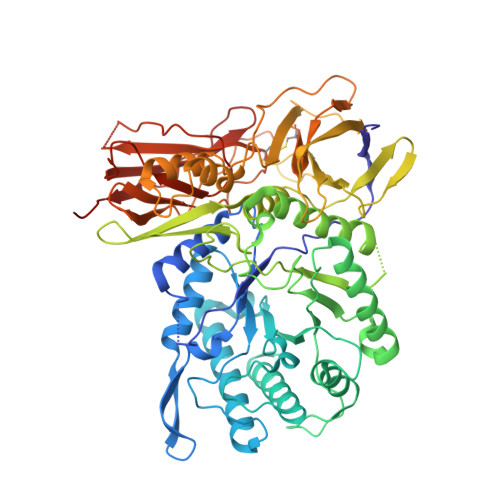
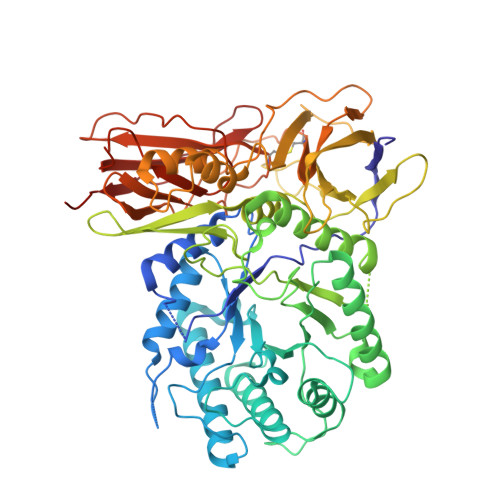
Cyclophellitol aziridines are potent irreversible inhibitors of retaining glycosidases and versatile intermediates in the synthesis of activity-based glycosidase probes (ABPs). Direct 3-amino-2-(trifluoromethyl)quinazolin-4(3H)-one-mediated aziridination of l-ido-configured cyclohexene has enabled the synthesis of new covalent inhibitors and ABPs of α-l-iduronidase, deficiency of which underlies the lysosomal storage disorder mucopolysaccharidosis type I (MPS I). The iduronidase ABPs react covalently and irreversibly in an activity-based manner with human recombinant α-l-iduronidase (rIDUA, Aldurazyme ® ). The structures of IDUA when complexed with the inhibitors in a non-covalent transition state mimicking form and a covalent enzyme-bound form provide insights into its conformational itinerary. Inhibitors 1-3 adopt a half-chair conformation in solution ( 4 H 3 and 3 H 4 ), as predicted by DFT calculations, which is different from the conformation of the Michaelis complex observed by crystallographic studies. Consequently, 1-3 may need to overcome an energy barrier in order to switch from the 4 H 3 conformation to the transition state ( 2, 5 B) binding conformation before reacting and adopting a covalent 5 S 1 conformation. rIDUA can be labeled with fluorescent Cy5 ABP 2, which allows monitoring of the delivery of therapeutic recombinant enzyme to lysosomes, as is intended in enzyme replacement therapy for the treatment of MPS I patients.
Organizational Affiliation:
Department of Bio-organic Synthesis, Leiden Institute of Chemistry, Leiden University, Einsteinweg 55, 2333 CC, Leiden, The Netherlands.