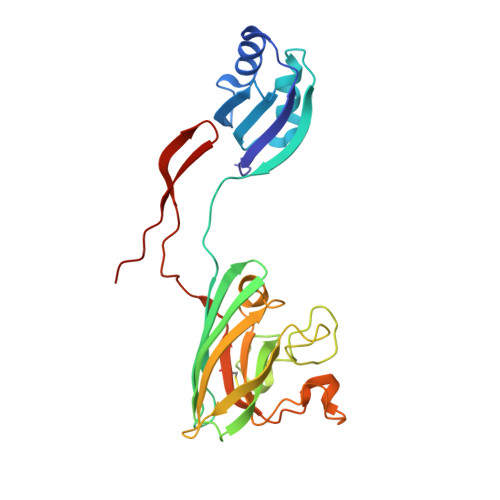
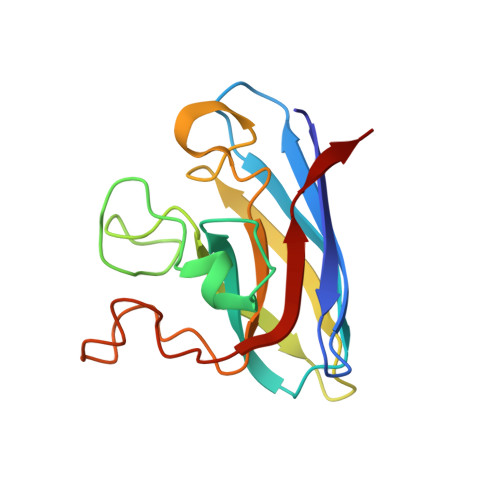
Molecular recognition and maturation of SOD1 by its evolutionarily destabilised cognate chaperone hCCS.
Sala, F.A., Wright, G.S.A., Antonyuk, S.V., Garratt, R.C., Hasnain, S.S.(2019) PLoS Biol 17: e3000141-e3000141
- PubMed: 30735496
- DOI: https://doi.org/10.1371/journal.pbio.3000141
- Primary Citation of Related Structures:
6FN8, 6FOI, 6FOL, 6FON, 6FP6 - PubMed Abstract:
Superoxide dismutase-1 (SOD1) maturation comprises a string of posttranslational modifications which transform the nascent peptide into a stable and active enzyme. The successive folding, metal ion binding, and disulphide acquisition steps in this pathway can be catalysed through a direct interaction with the copper chaperone for SOD1 (CCS). This process confers enzymatic activity and reduces access to noncanonical, aggregation-prone states. Here, we present the functional mechanisms of human copper chaperone for SOD1 (hCCS)-catalysed SOD1 activation based on crystal structures of reaction precursors, intermediates, and products. Molecular recognition of immature SOD1 by hCCS is driven by several interface interactions, which provide an extended surface upon which SOD1 folds. Induced-fit complexation is reliant on the structural plasticity of the immature SOD1 disulphide sub-loop, a characteristic which contributes to misfolding and aggregation in neurodegenerative disease. Complexation specifically stabilises the SOD1 disulphide sub-loop, priming it and the active site for copper transfer, while delaying disulphide formation and complex dissociation. Critically, a single destabilising amino acid substitution within the hCCS interface reduces hCCS homodimer affinity, creating a pool of hCCS available to interact with immature SOD1. hCCS substrate specificity, segregation between solvent and biological membranes, and interaction transience are direct results of this substitution. In this way, hCCS-catalysed SOD1 maturation is finessed to minimise copper wastage and reduce production of potentially toxic SOD1 species.
Organizational Affiliation:
Molecular Biophysics Group, Institute of Integrative Biology, Faculty of Health and Life Sciences, University of Liverpool, Liverpool, United Kingdom.